INTRODUCTION
Geldanamycin (GDM) is a benzoquinone ansamycin antibiotic. Recently, GDM was isolated from Streptomyces zerumbet W14 (Taechowisan et al., 2019). It interacts with heat shock protein 90 (Hsp90), which inhibits the growth of cancers (Mimnaugh et al., 1996; Whitesell et al., 1994). However, the utilization management of this compound has been limited by its poor water solubility and hepatotoxicity (Fukuyo et al., 2009; Supko et al., 1995). Therefore, the development of GDM derivatives with increased pharmacokinetic properties has been attempted. Various synthetic GDM derivatives that will generate new types of Hsp90 inhibitors with weak toxicity and high efficiency have been sought (Kitson et al., 2013; Lin et al., 2015; Modi et al., 2011; Tian et al., 2004; Wrona et al., 2010). Several GDM derivatives have been synthesized, for example, 17-N-allylamino-17-demethoxygeldanamycin, and 17-dimethylamino ethylamino-17-demethoxygeldanamycin; however, their water solubility was limited (Smith et al., 2005). Recently, tryptamine-geldanamycin hybrids have been synthesized. These compounds showed moderate cytotoxic activity against normal cells and strong cytotoxic activity against various cancer cell lines. Their water solubility was increased above that of GDM (Taechowisan et al., 2020).
Dopamine is a neurotransmitter responsible for transmitting signals between nerve cells. It is used in the treatment of severe hypotension, bradycardia, circulatory shock, and cardiac arrest (Bhatt-Mehta and Nahata, 1989). Dopamine is polar a covalent compound that is soluble in polar molecules such as water. In this study, it is a useful tool for the development of dopamine-geldanamycin hybrids (DGH) with improved solubility and biological activities. The C17-methoxy group of the GDM molecule can allow the introduction of various nucleophiles. Thus, GDM has been a precursor for synthesizing its derivatives (Modi et al., 2011; Tian et al., 2004; Wrona et al., 2010). Furthermore, another report showed that, compared with GDM, some of the GDM derivatives exhibited greater activity against myeloma cells (Jurczyszyn et al., 2014; Mielczarek-Lewandowska et al., 2019).
In this study, the novel DGH was synthesized. Molecular docking was performed to determine the hydrogen-bonding interaction, the binding energy, and the orientation of inhibitors bound in the active site of Hsp90. The ADMET of the compounds was predicted via in silico methods. Their anticancer activity was then evaluated on three cancer cell lines, MDA-MB-231, HeLa, and human hepatocellular carcinoma (HepG2) cells, using an MTT colorimetric assay. The water solubility was also carried out.
MATERIALS AND METHODS
Cultivation of actinomycetes and product isolation
Streptomyces zerumbet W14 was obtained from Zingiber zerumbet (L.) Smith using the surface-sterilization technique (Taechowisan et al., 2019). The bacterium was cultured on ISP-2 agar at 30oC for 14 days. The initial steps of antibiotic isolation and purification were as previously described (Taechowisan et al., 2020). The isolated compound was subjected to investigation by NMR spectroscopy. The spectral data for this compound identified it as geldanamycin (C29H40N2O9) (1).
Chemical reagents and materials
The chemicals were purchased from Sigma-Aldrich (Darmstadt, Germany), Tokyo Chemical Industry (Tokyo, Japan), and Fluka Chemical (Buchs, Switzerland) Companies. The 1H and 13C NMR spectra were recorded with a Bruker Avance 300 spectrometer (Bruker, MA). Mass spectra were determined with a micrOTOF (Bruker, MA). Melting points were measured with a Stuart Scientific SMP 2 (Cole-Parmer Ltd., Staffordshire, UK) and are uncorrected. The reaction was monitored by TLC, performed on aluminum sheets precoated with silica gel 60 (Darmstadt, Germany). Column chromatography was carried out using a Merck Kieselgel 60 column chromatography (Darmstadt, Germany).
Synthesis of DGH
The synthesis of DGH has been described by Taechowisan et al. (2021). The spectral data of the synthesized compounds are described as follows:
(4E,6Z,8S,9S,10E,12S,13R,14S,16R)-19-((3,4-dihydroxyphenethyl)amino)-13-hydroxy-8,14-dimethoxy-4,10,12,16-tetramethyl-3,20,22-trioxo-2-azabicyclo[16.3.1]docosa-1(21),4,6,10,18-pentaen-9-yl carbamate (2): 0.0167 g (21% yield using 1:2 hexane:EtOAc as a mobile phase); m.p. 114oC–116oC; 1H-NMR (CDCl3, 300 MHz) δ 0.93 (d, J = 6.2 Hz, 3H, CH3), 1.00 (d, J = 6.9 Hz, 3H, CH3), 1.60–1.78 (m, 3H, hydrocarbon), 1.80 (s, 3H, CH3), 2.01 (s, 3H, CH3), 2.35–2.43 (m, 1H, CH), 2.62–2.83 (m, 4H, hydrocarbon), 3.27 (s, 3H, OCH3), 3.36 (s, 3H, OCH3), 3.46 (m, 1H, CH), 3.59–3.73 (m, 3H, hydrocarbon), 4.33 (d, J = 9.7 Hz, 1H, CH), 4.97 (br, 2H, NH2), 5.18 (s, 1H, CH), 5.83–5.90 (m, 2H, 2CH), 6.41 (br, 1H, NH), 6.54–6.61 (m, 2H, 2CH), 6.73 (d, J = 1.8 Hz, 1H, ArH), 6.81 (d, J = 8.0 Hz, 1H, ArH), 6.94 (d, J = 11.5 Hz, 1H, CH), 7.21 (s, 1H, CH), 9.18 (br, 1H, NH); 13C-NMR (CDCl3, 75 MHz) δ 12.5, 12.6, 12.8, 22.8, 28.6, 29.7, 32.3, 34.3, 35.0, 35.2, 47.0, 56.7, 57.1, 72.8, 81.1, 81.4, 81.9, 108.5, 108.7, 115.6, 115.7, 120.8, 126.6, 127.0, 129.6, 132.8, 133.6, 134.9, 141.3, 143.1, 144.3, 144.9, 156.3, 168.5, 180.4, 183.9; HMS calculated for C36H47N3O10 (M+Na)+ 704.3154, found 704.3157.
(4E,6Z,8S,9S,10E,12S,13R,14S,16R)-19-((3,4-dimethoxyphenethyl)amino)-13-hydroxy-8,14-dimethoxy-4,10,12,16-tetramethyl-3,20,22-trioxo-2-azabicyclo[16.3.1]docosa-1(21),4,6,10,18-pentaen-9-yl carbamate (3): 0.0508 g (50% yield using 1:1 Hexane:EtOAc as a mobile phase); m.p. 127oC–129oC; 1H-NMR (CDCl3, 300 MHz) δ 0.95 (d, J = 6.4 Hz, 3H, CH3), 1.00 (d, J = 6.9 Hz, 3H, CH3), 1.62–1.78 (m, 3H, hydrocarbon), 1.80 (s, 3H, CH3), 2.02 (s, 3H, CH3), 2.39–2.46 (m, 1H, CH), 2.67–2.93 (m, 4H, hydrocarbon), 3.27 (s, 3H, OCH3), 3.36 (s, 3H, OCH3), 3.45 (m, 1H, CH), 3.57 (dd, J = 1.8, 9.0 Hz, 1H, CH), 3.62–3.85 (m, 1H, CH), 3.87 (s, 3H, OCH3), 3.89 (s, 3H, OCH3), 4.31 (d, J = 9.8 Hz, 1H, CH), 4.86 (br, 2H, NH2), 5.19 (s, 1H, CH), 5.83–5.92 (m, 2H, 2CH), 6.38 (br, 1H, NH), 6.58 (t, J = 11.5 Hz, 1H, CH), 6.71–6.91 (m, 3H, 3ArH), 6.95 (d, J = 11.4 Hz, 1H, CH), 7.26 (s, 1H, CH), 9.17 (br, 1H, NH); 13C-NMR (CDCl3, 75 MHz) δ 12.3, 12.6, 12.8, 22.9, 28.5, 32.3, 34.4, 35.1, 35.5, 47.1, 55.9, 56.7, 57.1, 72.65, 81.2, 81.4, 81.6, 108.6, 108.7, 111.6, 111.7, 120.7, 126.5, 126.9, 129.6, 132.7, 133.7, 134.9, 135.8, 141.3, 144.7, 148.2, 149.3, 156.0, 168.4, 180.7, 183.8; HMS calculated for C38H51N3O10 (M+Na)+ 732.3467, found 732.3469.
(4E,6Z,8S,9S,10E,12S,13R,14S,16R)-19-((3,4-bis(benzyloxy)phenethyl)amino)-13-hydroxy-8,14-dimethoxy-4,10,12,16-tetramethyl-3,20,22-trioxo-2-azabicyclo[16.3.1]docosa-1(21),4,6,10,18-pentaen-9-yl carbamate (4): 0.0501 g (14% yield using 5:2 CH2Cl2:EtOAc as a mobile phase); m.p. 92oC–94oC; 1H-NMR (CDCl3, 300 MHz) δ 0.93 (d, J = 6.4 Hz, 3H, CH3), 1.00 (d, J = 6.9 Hz, 3H, CH3), 1.62–1.78 (m, 3H, hydrocarbon), 1.79 (s, 3H, CH3), 2.02 (s, 3H, CH3), 2.34–2.42 (m, 1H, CH), 2.64–2.77 (m, 1H, CH), 2.84 (t, J = 6.9 Hz, 2H, CH2), 3.26 (s, 3H, OCH3), 3.36 (s, 3H, OCH3), 3.43–3.46 (m, 1H, CH), 3.58 (dd, J = 1.7, 8.9 Hz, 1H, CH), 3.62–3.82 (m, 2H, CH2), 4.31 (d, J = 9.9 Hz, 1H, CH), 4.96 (br, 2H, NH2), 5.14 (s, 2H, OCH2), 5.16 (s, 2H, OCH2), 5.18 (s, 1H, CH), 5.82–5.91 (m, 2H, 2CH), 6.32 (br, 1H, NH), 6.58 (t, J = 11.5 Hz, 1H, CH), 6.73 (dd, J = 2.0, 8.1 Hz, 1H, ArH), 6.79 (d, J = 2.0 Hz, 1H, ArH), 6.91 (d, J = 8.2 Hz, 1H, ArH), 6.95 (d, J = 10.8 Hz, 1H, CH), 7.26 (s, 1H, CH), 7.29–7.40 (m, 6H, 6ArH), 7.40–7.55 (m, 4H, 4ArH), 9.17 (br, 1H, NH); 13C-NMR (CDCl3, 75 MHz) δ 12.3, 12.6, 12.7, 22.9, 28.5, 32.3, 34.4, 35.0, 35.4, 46.9, 56.7, 57.1, 71.4, 71.5, 72.6, 81.2, 81.4, 81.6, 108.6, 108.7, 115.4, 115.6, 121.6, 126.5, 126.9, 127.3, 127.8, 128.5, 130.4, 132.7, 133.7, 134.9, 135.8, 137.1, 137.2, 141.3, 144.7, 148.2, 149.3, 156.1, 168.3, 180.6, 183.7; HMS calculated for C50H59N3O10 (M+Na)+ 884.4093, found 884.4094.
(4E,6Z,8S,9S,10E,12S,13R,14S,16R)-19-((3,4-bis((2-bromobenzyl)oxy)phenethyl) amino)-13-hydroxy-8,14-dimethoxy-4,10,12,16-tetramethyl-3,20,22-trioxo-2-azabicyclo[16.3.1]do-cosa-1(21),4,6,10,18-pentaen-9-yl carbamate (5): 0.0757 g (53% yield using 2:1 CH2Cl2:EtOAc as a mobile phase); m.p. 174oC–176oC; 1H-NMR (CDCl3, 300 MHz) 0.94 (d, J = 6.4 Hz, 3H, CH3), 1.00 (d, J = 6.9 Hz, 3H, CH3), 1.62–1.79 (m, 3H, hydrocarbon), 1.80 (s, 3H, CH3), 2.02 (s, 3H, CH3), 2.36–2.44 (m, 1H, CH), 2.65–2.80 (m, 1H, CH), 2.88 (t, J = 6.9 Hz, 2H, CH2), 3.26 (s, 3H, OCH3), 3.36 (s, 3H, OCH3), 3.43–3.46 (m, 1H, CH), 3.57 (dd, J = 1.4, 8.6 Hz, 1H, CH), 3.60–3.85 (m, 2H, CH2), 4.31 (d, J = 9.9 Hz, 1H, CH), 5.06 (br, 2H, NH2), 5.18 (s, 1H, CH), 5.20 (s, 2H, OCH2), 5.22 (s, 2H, OCH2), 5.82–5.91 (m, 2H, 2CH), 6.33 (br, 1H, NH), 6.58 (t, J = 11.6 Hz, 1H, CH), 6.77–6.86 (m, 2H, 2ArH), 6.93 (d, J = 8.1 Hz, 1H, ArH), 6.95 (d, J = 11.6 Hz, 1H, CH), 7.17 (td, J = 0.5, 7.4 Hz, 2H, 2ArH), 7.26 (s, 1H, CH), 7.31 (td, J = 0.6, 7.5 Hz, 2H, 2ArH), 7.56 (d, J = 7.9 Hz, 2H, ArH), 7.61 (ddd, J = 1.8, 2.3, 7.7 Hz, 2H, 2ArH), 9.17 (br, 1H, NH); 13C-NMR (CDCl3, 75 MHz) δ 12.3, 12.5, 12.7, 22.9, 28.4, 32.3, 34.4, 35.0, 35.3, 46.8, 56.7, 57.0, 70.6, 72.5, 81.2, 81.4, 81.5, 108.6, 108.7, 115.1, 115.2, 121.9, 122.0, 122.1, 126.5, 126.9, 127.5, 127.6, 128.8, 128.9, 129.1, 129.2, 130.7, 130.8, 132.4, 132.8, 133.6, 134.9, 135.8, 136.3, 136.4, 141.2, 144.7, 147.7, 147.8, 148.8, 148.9, 156.2, 168.3, 180.6, 183.6; HMS calculated for C50H57Br2N3O10 (M+Na)+ 1040.2303, found 1040.2323.
(4E,6Z,8S,9S,10E,12S,13R, 14S,16R)-19-((3,4-dibutoxyphenethyl)amino)-13-hydroxy-8,14-dimethoxy-4,10,12,16-tetramethyl-3,20,22-trioxo-2-azabicyclo [16.3.1]docosa-1(21),4,6,10,18-pentaen-9-yl carbamate (6): 0.0097 g (25.8% yield using 10:7 CH2Cl2:EtOAc as the mobile phase); m.p. 178oC–181oC; 1H-NMR (CDCl3, 300 MHz) δ 0.98–1.01 (m, 12H, 4CH3), 1.41–1.60 (m, 4H, 2CH2), 1.61–1.89 (m, 10H, hydrocarbon), 2.02 (s, 3H, CH3), 2.38–2.46 (m, 1H, CH), 2.65–2.80 (m, 2H, hydrocarbon), 2.88 (t, J = 7.5 Hz, 2H, CH2), 3.27 (s, 3H, OCH3), 3.36 (s, 3H, OCH3), 3.45 (m, 1H, CH), 3.57 (dd, J = 1.8, 8.9 Hz, 1H, CH), 3.62–3.87 (m, 2H, CH2), 3.99 (td, J = 3.9, 6.6 Hz, 4H, 2CH2), 4.31 (d, J = 9.9 Hz, 1H, CH), 4.84 (br, 2H, NH2), 5.19 (s, 1H, CH), 5.83–5.92 (m, 2H, 2CH), 6.37 (br, 1H, NH), 6.58 (t, J = 11.6 Hz, 1H, CH), 6.72–6.75 (m, 2H, 2ArH), 6.85 (d, J = 8.67 Hz, 1H, ArH), 6.95 (d, J = 11.7 Hz, 1H, CH), 7.26 (s, 1H, CH), 9.17 (br, 1H, NH); 13C-NMR (CDCl3, 75 MHz) δ 12.3, 12.6, 12.7, 13.8, 19.2, 22.9, 28.5, 31.3, 32.3, 34.4, 35.1, 35.5, 47.1, 56.7, 57.1, 69.1, 72.6, 81.2, 81.5, 81.7, 108.6, 108.7, 114.3, 114.4, 120.9, 126.5, 126.9, 129.7, 132.7, 133.8, 135.0, 135.8, 141.3, 144.8, 148.4, 149.5, 156.0, 168.4, 180.6, 183.7; HMS calculated for C44H63N3O10 (M+Na)+ 816.4406, found 816.4402.
The water solubility of DGH was carried out by comparison with GDM as previously described (Taechowisan et al., 2020).
MTT assay for cell viability
Two normal cell lines [African green monkey kidney cells (Vero); Rhesus monkey kidney cells (LLC-MK2)] and three cancer cell lines [human breast carcinoma cells (MDA-MB-231]; human hepatocellular carcinoma cells (HepG2); and human cervical carcinoma cells (HeLa) (from the Korean Cell Line Bank)) were grown in DMEM medium supplemented with 10% in a 37°C, 5% CO2 humidified incubator. Cytotoxicity assay was carried out as described in a previous publication (Taechowisan et al., 2020).
Molecular docking analysis
The two-dimensional structures of geldanamycin and DGH were drawn and converted to SMILES strings with ChemDraw software (http://cambridgesoft.com) and the SMILES Translator and Structure Generator (https://cactus.nci.nih.gov/translate/), respectively. The energies of these compounds were optimized and converted to #D format, saved as Protein Data Bank (PDB) files using UCSF Chimera v.1.14 (University of California, CA), and further used for molecular docking analysis.
The 3D structure of Hsp90 with the cocrystallized geldanamycin (1YET) was retrieved from the PDB and chosen for molecular docking studies. The 3D structure of 1YET is taken for docking geldanamycin and DGH to obtain the predictions of ligand binding. The water molecules were taken out from its structure using Discovery Studio software (Accelrys Inc.). Docking simulations were undertaken with Hsp90 as the target (1YET) and geldanamycin, 17-DMAG, and DGH as the ligands using AutoDock Vina to predict the ligand-binding sites on Hsp90. Docking analysis was performed as detailed by Taechowisan et al. (2021).
Prediction of ADMET by computational analysis
The computational prediction of the compounds was performed using two types of online software SwissADME (http://swissadme.ch) and Pre-ADMET (https://preadmet.bmdrc.kr) to obtain relative results for pharmacokinetic profile (absorption, distribution, metabolism, excretion, and toxicity; ADMET) of the molecules. The ADMET profiles of DGH were analyzed in comparison with geldanamycin.
Statistical analysis
Each result represents the means ± standard deviation of three experiments. SPSS v.16.0 (SPSS Inc., Chicago, IL) software was used for the data analysis. Comparisons between the two groups were analyzed using the two-tailed Dunnett’s t-tests treated compound 1 as a control group. A p-value < 0.05 was considered to indicate statistical significance.
RESULTS
In the present work, a series of DGH were synthesized via nucleophilic substitution of GDM (1) as described in Methods section (Schemes 1 and 2).
The solubility in water of compound 1 was found to be 0.152 mM (Table 1). In contrast, the solubility of DGH in water was between 0.386 mM and 5.464 mM, approximately 2.53–35.94 times better than that of GDM. These data suggest that the conjugation of a dopamine moiety to GDM at the C17-position greatly enhanced their solubility in water.
GDM and DGH were also assessed for cytotoxicity activity against two normal cell lines (Vero and LLC-MK2 cells) and three cancer cell lines (MDA-MB-231, HepG2, and HeLa) using the MTT assay. All DGH showed weak cytotoxicity activity toward Vero and LLC-MK2 cells with IC50 values in the range of 104.52–496.31 μg/ml (Table 2). The results show that DGH has low toxicity to normal cells compared with GDM and doxorubicin. All DGH showed less cytotoxicity activity toward HeLa cells than geldanamycin. Compounds 2, 3, and 6 were also active against MDA-MB231 cells with IC50 values of 41.88, 52.12, and 70.93 μg/ml, respectively. Amongst these compounds, only compound 6 was activity against HeLa cells with IC50 values of 89.38 μg/ml. Compounds 4 and 5 were not active on cytotoxicity against both normal cells and cancer cells. In addition, the therapeutic index of DGH is greater than those of GDM, informing that DGH has safety as the concentration required to cause toxicity. It suggested that some of them (compounds 2, 3, and 6) were toxic to some cancer cells. Therefore, they could be the potential drug candidate against some cancers.
 | Scheme 1. Syntheses of dopamine derivatives 10a–10d. [Click here to view] |
 | Scheme 2. Syntheses of 17-demethoxygeldanamycin derivatives 2–6. [Click here to view] |
Hsp90; PBD ID: 1YET was selected for molecular docking studies. Comparative docking of 1YET with 17-DMAG, GDM, and DGH was carried out to provide evidence in support of their inhibitory effects on Hsp90 interactions. The results of docking studies (as shown in Table 3) revealed that 17-DMAG participated in interactions through five hydrogen bonds with Ser52, Asp54, Lys58, Asp93, and Phe138 to the N-terminal ATP-binding pocket of Hsp90, with a binding energy of −136.55 kcal/mol. Geldanamycin bridged four hydrogen bonds with Ser52, Lys58, Asp93, and Phe138 to the N-terminal ATP-binding pocket of Hsp90, with a binding energy of −133.06 kcal/mol. Compound 2 formed six hydrogen bonds with Glu47, Asn106 (three positions), Gly137, and Phe138 to the N-terminal ATP-binding pocket, with a binding energy of −101.06 kcal/mol. Compound 3 formed three hydrogen bonds with Lys58, Gly137, and Phe138 to the N-terminal ATP-binding pocket, with a binding energy of −98.54 kcal/mol. Compound 4 formed two hydrogen bonds with Gln212 and Ile218 to the N-terminal part of Hsp90 (outside active site), with a binding energy of −108.12 kcal/mol. Compound 5 did not form a hydrogen bond to any part of the Hsp90 molecule. Compound 6 formed three hydrogen bonds with Glu47, Lys112, and Gly137 to the N-terminal ATP-binding pocket of Hsp90, with a binding energy of −97.03 kcal/mol. Figure 1 displays the 2D interactions of the compounds and Hsp90 obtained from molecular docking using LIGPLOT. As can be seen, 17-DMAG formed the highest number of hydrogen bonds (Fig. 1a). To display the docking results, compound 4 participated in interactions outside the active site of the N-terminal part of Hsp90 and compound 5 did not form a hydrogen bond to any part of the Hsp90 molecule; they were carried out using the BIOVIA Discovery Studio as shown in Figure 2e and f, respectively. Figure 3 shows the hydrogen bond acceptor and hydrogen bond donors and how the ligand is set inside the cavity. The green patches present denote the hydrogen bond acceptor and the pink patches represent the hydrogen bond donor and they show how the ligand is present inside the cavity. As can be seen, compound 5 did not participate in the hydrogen bond acceptor or donor to any part of the Hsp90 molecule (Fig. 3f).
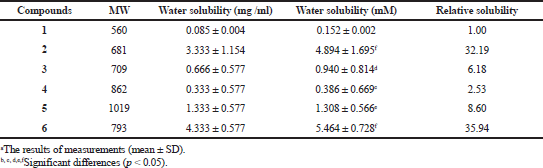 | Table 1. The solubility of geldanamycin (1) and synthetic compounds (2 to 6). [Click here to view] |
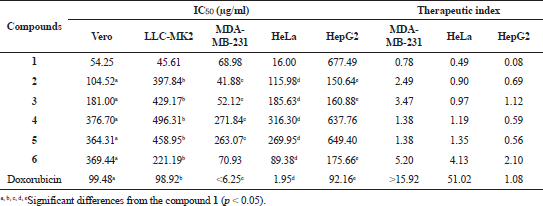 | Table 2. Cytotoxicity activity (IC50) of geldanamycin (1) and synthetic compounds (2 to 6). [Click here to view] |
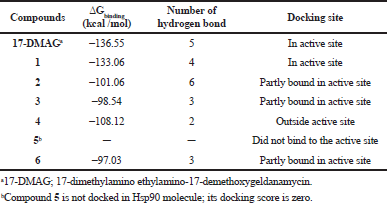 | Table 3. Geldanamycin (1) and its derivatives (2 to 6) to inhibit Hsp90 protein based on the molecular docking simulation. [Click here to view] |
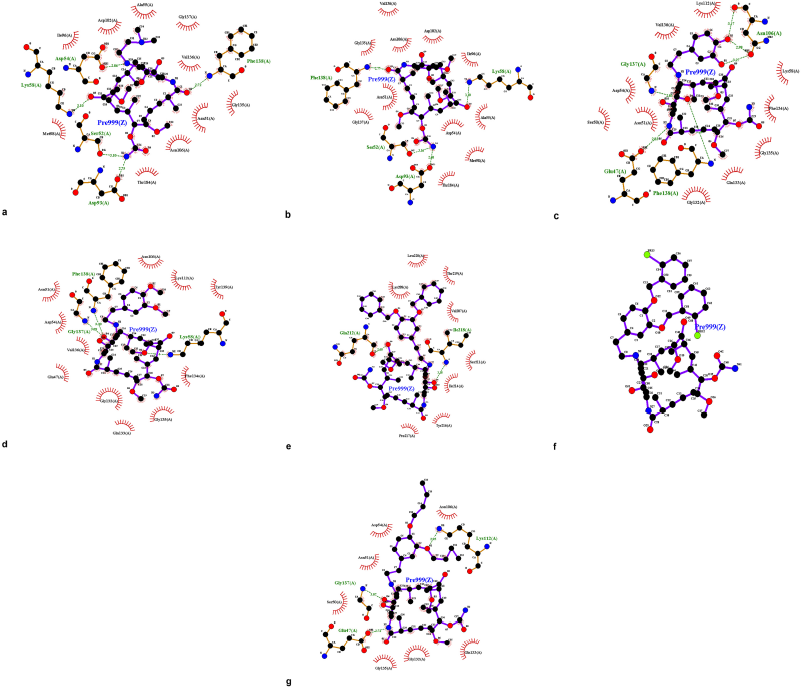 | Figure 1. LIGPLOT diagram showing hydrogen-bonding interactions (the green dashed lines) and hydrophobic contacts (the red crescents with the bristles) for the ligand: 17-DMAG (a) and compounds 1 to 6 (b to g) with Hsp90 molecule, respectively. [Click here to view] |
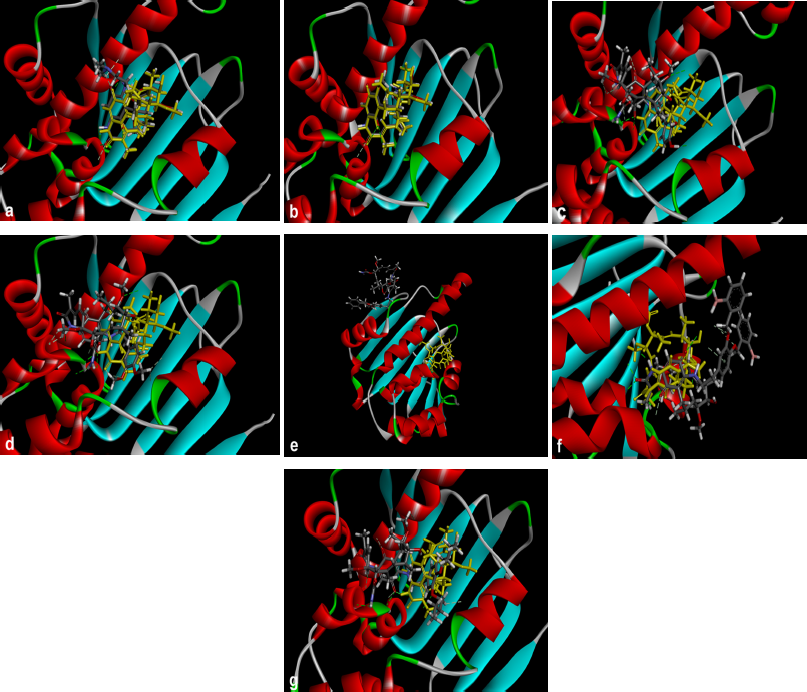 | Figure 2. Crystal structure superimposed on Hsp90 (1YET) docked 17-DMAG (a) and compounds 1 to 6 (b to g) with Hsp90 molecule, respectively. Compound 5 was not docked in Hsp90 molecule (f). The brown molecule was represented by geldanamycin. [Click here to view] |
The predicted ADMET profiles of GDM and DGH are presented in Table 4. The compounds have high polarity (hydrophilic); the TPSA values are greater than 140. Compounds 1, 2, 3, and 6 have ideal lipophilicity (average of Log Po/w ≤ 5). This result suggested that these compounds have good absorption and permeation. The predicted values of Caco2-cell permeability are ≥ 0.90; therefore, these compounds have high Caco2-cell permeability and are the ease of absorption. These compounds have good intestinal absorption, because the predicted values of human intestinal absorption are greater than 30%. These compounds are predicted to have high skin permeability (the log Kp < −2.5). The compounds are predicted to be substrates of P-glycoprotein; they may be excreted from cells by P-glycoprotein. However, compounds 1, 2, and 3 are also predicted not to be a P-glycoprotein inhibitor. The compounds 1, 2, 3, and 4 are predicted to moderately cross the blood-brain barrier (−1 < logBB < 0.3), but compounds 5 and 6 easily crossed (logBB > 0.3). These compounds are substrates for CYP3A4, but not substrates for CYP2D6. They are also predicted to be CYP3A4 inhibitor. Therefore, they may be metabolized in the liver. They also suggest that these compounds have not mutagenic activity by AMES test, but they may inhibit the hERG channel and may cause cardiotoxicity. The predicted results indicate that the ADMET properties of DGH are similar to those of GDM. However, they were a cytochrome substrate and inhibitor and hERG inhibitor, which may cause hepatotoxicity and cardiotoxicity, respectively. This should be taken with precautions.
 | Figure 3. Hydrogen-bonding interactions of 17-DMAG (a), compound 1 (b), compounds 2, 3, and 6 (c, d, and g) molecules in the part of the active pocket site, compound 4 molecule at the outside of the active pocket site (e), and compound 5 molecule did not bind to any part of Hsp90 (f), respectively. [Click here to view] |
DISCUSSION
GDM was the first benzoquinone ansamycin antibiotic generated by Streptomyces hygroscopicus var. geldanus which was discovered in 1970 (Johnson et al., 2010). Recently, it was also isolated from S. zerumbet W14 (Taechowisan et al., 2019). It specifically targets and deactivates Hsp90. For the past decade, Hsp90 has been an interesting target molecule for anticancer treatment, because it plays a role in protein functions and cell growth (Chatterjee and Burns, 2017). Although GDM is a potent cell growth inhibitor, however, it suffers from severe hepatotoxicity and insolubility in water (Fukuyo et al., 2009; Supko et al., 1995). Therefore, the structure modification of this compound to reduce hepatotoxicity and increase water solubility should be carried out. Some of geldanamycin derivatives are in clinical trials, for example, 17-allylamine-17-demethoxygeldanamycin (17-AAG) (Banerji et al., 2003; Goetz et al., 2005; Heath et al., 2008; Ramanathan et al., 2005; Solit et al., 2008), 17-(dimethylaminoethylamino)-17-demethoxygeldanamycin (17-DMAG) (Grem et al., 2005), and 17-allylamino-17-demethoxygeldanamycin hydroquinone hydrochloride (Hanson and Vesole, 2009; Sydor et al., 2006). However, the toxicity of these compounds was unfavorable, as they caused both cardiac and liver toxicity (Lancet et al., 2006; Sequist and Janne, 2007). Meanwhile, novel geldanamycin derivatives have been synthesized which are excellent tools for exploring their biological activities.
In this study, five new geldanamycin derivatives, compounds 2–6, were synthesized by nucleophilic substitution of GDM (1). Position 17 of geldanamycin has been introduced for the synthesis of geldanamycin compounds because its methoxy group is readily displaced by a nucleophile, providing a convenient entry into 17-substituted-17-demethoxygeldanamycin compounds by the SN2 reaction mechanism (Hamlin et al., 2018). By the molecular docking studies, compounds 2, 3, and 6 partly bound in the active site of the N-terminal ATP-binding pocket of Hsp90. The binding mode of these compounds with Hsp90 (1YET) using the 3D structure has been predicted to understand the protein-ligand contacts and their interaction. The docking results of these compounds were compared along with 17-DMAG (control compound) in the active site of 1YET. These DGH showed a similar pose as that of 17-DMAG and GDM in the binding pocket. The binding energy of these DGH is in the range of −97.03 to −101.06 kcal/mol, in comparison to 17-DMAG and GDM with a binding energy of −136.55 and −133.06 kcal/mol, respectively. The LIGPLOT showed that DGH formed hydrogen bonding with most similar to that observed in 17-DMAG, and also they showed highly conserved hydrophobic interactions with the same amino acid residues. These interactions occurred on the benzoquinone moiety of geldanamycin molecule. Similar to the findings of previous studies, the molecular docking in the present study has shown that the pocket of Hsp90 is composed of a mixture of polar, charged, and hydrophobic amino acids. These amino acids include Asn51, Asp54, Ala55, Lys58, Asp93, Met98, Asn106, Leu107, Lys112, Gly135, Phe138, and Thr184. Therefore, the bottom of the pocket becomes increasingly hydrophobic; it retains one polar residue and one charged residue at the deepest portion as Thr184 and Asp93, respectively (Abbasi et al., 2017; Stebbins et al., 1997; Teo et al., 2015). The ADMET parameters of the compounds were studied by two types of online software. The correlation of structure and toxicity of these compounds was preliminarily evaluated. These compounds have good absorption and permeation. They are the substrate of P-glycoprotein, so they may be excreted from cells to the body because they also have the potential for cytochrome P450. They are predicted to be metabolized in the liver which may have hepatotoxicity by redox metabolism and glutathione adduct formation (Cysyk et al., 2006; Guo et al., 2008). Published reports suggest that the benzoquinone moiety in the geldanamycin molecule is responsible for hepatotoxicity (Cysyk et al., 2006; Guo et al., 2008).
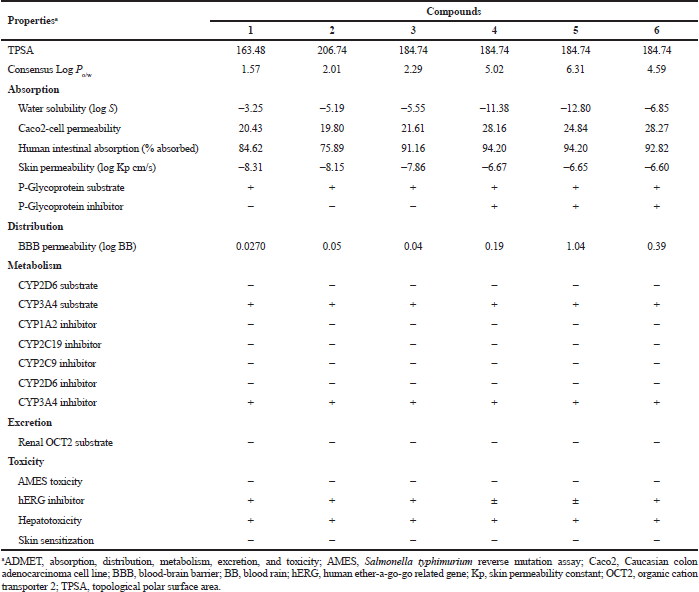 | Table 4. Predicted ADMET properties of geldanamycin (1) and synthetic compounds (2 to 6). [Click here to view] |
In the present study, the experimental basis was carried out; compounds 2, 3, and 6 showed a greater increase in water solubility and also showed less cytotoxicity than GDM in the normal cell lines and presented greater toxicity to some cancer cell lines. This compound should increase the likelihood of use in future studies. However, compound 4 bound outside the active side and compound 5 did not bind to any part of Hsp90. They were not active on cytotoxicity against both normal cells and cancer cells. The findings confirm that the N-terminal ATP-binding domain is essential for the function of Hsp90 as described previously (Stebbins et al. 1997). Doxorubicin is a broad-spectrum chemotherapeutic drug used to treat cancers. In this study, it was used as the positive control. Doxorubicin exhibited greater cytotoxicity against MDA-MB231 and HeLa cells than GDM and DGH. The mechanisms of anticancer activity of doxorubicin were (i) DNA intercalation and topoisomerase-II disruption; (ii) free radical generation and damage to proteins, DNA, and cellular membranes, which were different from geldanamycin and derivatives (Gewirtz, 1999); in addition, the side effect for the use of doxorubicin was cardiotoxicity and doxorubicin resistance was also a problem (Thorn et al., 2011; Weiss, 1992). Therefore, the development of Hsp90 inhibitors may become a universal chemotherapeutic of cancers.
CONCLUSION
In summary, novel DGH with anticancer activity, enhanced water solubility, and low toxicity was presented in this study, in comparison with GDM. In particular, compounds 2, 3, and 6 showed anticancer activity against some cancer cells; moreover, the study findings revealed, through molecular docking and ADMET analysis, that the development of DGH improved the pharmacokinetic profiles of solubility, cytotoxicity, and anticancer activities. Therefore, Hsp90 could be an anticancer target, and some DGH could be considered as a candidate for anticancer agents.
ACKNOWLEDGMENT
The authors gratefully acknowledge support from the Faculty of Science, Silpakorn University, Nakhon Pathom, Thailand (Research Grant No. SRIF-JRG-2564-03).
AUTHOR CONTRIBUTIONS
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work. All the authors are eligible to be an author as per the international committee of medical journal editors (ICMJE) requirements/guidelines.
CONFLICTS OF INTEREST
The authors report no financial or any other conflicts of interest in this work.
ETHICAL APPROVALS
This study does not involve experiments on animals or human subjects.
PUBLISHER’S NOTE
This journal remains neutral with regard to jurisdictional claims in published institutional affiliation.
REFERENCES
Abbasi M, Sadeghi-Aliabadi H, Amanlou M. Prediction of new Hsp90 inhibitors based on 3,4-isoxazolediamide scaffold using QSAR study, molecular docking and molecular dynamic simulation. DARU, 2017; 25(1):17. CrossRef
Banerji U, Judson I, Workman P. The clinical applications of heat shock protein inhibitors in cancer – present and future. Curr Cancer Drug Targets, 2003; 3(5):385–90. CrossRef
Bhatt-Mehta V, Nahata MC. Dopamine and dobutamine in pediatric therapy. Pharmacotherapy, 1989; 9(5):303–14. CrossRef
Chatterjee S, Burns TF. Targeting heat shock proteins in cancer: a promising therapeutic approach. Int J Mol Sci, 2017; 18(9):1978. CrossRef
Cysyk RL, Parker RJ, Barchi JJ Jr, Steeg PS, Hartman NR, Strong JM. Reaction of geldanamycin and C17-substituted analogues with glutathione: product identifications and pharmacological implications. Chem Res Toxicol, 2006;19(3):376–81. CrossRef
Fukuyo Y, Hunt CR, Horikoshi N. Geldanamycin and its anti-cancer activities. Cancer Lett, 2009; 290(1):24–35. CrossRef
Gewirtz DA. A critical evaluation of the mechanisms of action proposed for the antitumor effects of the anthracycline antibiotics adriamycin and daunorubicin. Biochem Pharmacol, 1999; 57(7):727–41. CrossRef
Goetz MP, Toft D, Reid J, Ames M, Stensgard B, Safgren S, Adjei AA, Sloan J, Atherton P, Vasile V, Salazaar S, Adjei A, Croghan G, Erlichman C. Phase I trial of 17-allylamino-17-demethoxygeldanamycin in patients with advanced cancer. J Clin Oncol, 2005; 23(6):1078–87. CrossRef
Grem JL, Morrison G, Guo XD, Agnew E, Takimoto CH, Thomas R, Szabo E, Grochow L, Grollman F, Hamilton JM, Neckers L, Wilson RH. Phase I and pharmacologic study of 17-(allylamino)-17-demethoxygeldanamycin in adult patients with solid tumors. J Clin Oncol, 2005; 23(9):1885–93. CrossRef
Guo W, Reigan P, Siegel D, Ross D. Enzymatic reduction and glutathione conjugation of benzoquinone ansamycin Hsp90 inhibitors: relevance for toxicity and mechanism of action. Drug Metab Dispos, 2008; 36(10):2050–7. CrossRef
Hamlin TA, Swart M, Bickelhaupt FM. Nucleophilic substitution (SN2): dependence on nucleophile, leaving group, central atom, substituents, and solvent. Chem Phys Chem, 2018; 19(11):1315–30. CrossRef
Hanson BE, Vesole DH. Retaspimycin hydrochloride (IPI-504): a novel heat shock protein inhibitor as an anticancer agent. Exp Opin Investig Drugs, 2009; 18(9):1375–83. CrossRef
Heath EI, Hillman DW, Vaishampayan U, Sheng S, Sarkar F, Harper F, Gaskins M, Pitot HC, Tan W, Ivy SP, Pili R, Carducci MA, Erlichman C, Liu G. A phase II trial of 17-allylamino-17-demethoxygeldanamycin in patients with hormone-refractory metastatic prostate cancer. Clin Cancer Res, 2008; 14(23):7940–6. CrossRef
Johnson VA, Singh EK, Nazarova LA, Alexander LD, McAlpine SR. Macrocyclic inhibitors of hsp90. Curr Top Med Chem, 2010; 10(14):1380–402. CrossRef
Jurczyszyn A, Zebzda A, Czepiel J, Perucki W, Bazan-Socha S, Cibor D, Owczarek D, Majka M. Geldanamycin and its derivatives inhibit the growth of myeloma cells and reduce the expression of the MET receptor. J Cancer, 2014; 5(6):480–90. CrossRef
Kitson RR, Chang CH, Xiong R, Williams HE, Davis AL, Lewis W, Dehn DL, Siegel D, Roe SM, Prodromou C, Ross D. Synthesis of 19-substituted geldanamycins with altered conformations and their binding to heat shock protein hsp90. Nat Chem, 2013; 5(4):307–14. CrossRef
Lin Z, Peng R, Li Z, Wang Y, Lu C, Shen Y, Wang J, Shi, G. 17-ABAG, a novel geldanamycin derivative, inhibits LNCaP-cell proliferation though heat shock protein 90 inhibition. Int J Mol Med, 2015; 36(2):424–32. CrossRef
Lancet J, Baer MR, Gojo I, Burton M, Quinn M, Tighe SM, Bhalla K, Kersey K, Wells S, Zhong Z. Phase 1, pharmacokinetic (PK) and pharmacodynamic (PD) study of intravenous alvespimycin (KOS-1022) in patients with refractory hematological malignancies. Blood, ASH Annual Meeting, 2006; 108(11):1961. CrossRef
Mielczarek-Lewandowska A, Sztiller-Sikorska M, Osrodek M, Czyz M, Hartman ML. 17-Aminogeldanamycin selectively diminishes IRE1α-XBP1s pathway activity and cooperatively induces apoptosis with MEK1/2 and BRAFV600E inhibitors in melanoma cells of different genetic subtypes. Apoptosis, 2019; 24(7–8):596–611. CrossRef
Mimnaugh EG, Chavany C, Neckers L. Polyubiquitination and proteasomal degradation of the p185c-erbB-2 receptor protein-tyrosine kinase induced by geldanamycin. J Biol Chem, 1996; 271(37):22796–801. CrossRef
Modi S, Stopeck A, Linden H, Solit D, Chandarlapaty S, Rosen N, D'Andrea G, Dickler M, Moynahan ME, Sugarman S, Ma W. HSP90 inhibition is effective in breast cancer: a phase II trial of tanespimycin (17-AAG) plus trastuzumab in patients with HER2-positive metastatic breast cancer progressing on trastuzumab. Clin Cancer Res, 2011; 17(15):5132–9. CrossRef
Ramanathan RK, Trump DL, Eiseman JL, Belani CP, Agarwala SS, Zuhowski EG, Lan J, Potter DM, Ivy SP, Ramalingam S, Brufsky AM, Wong MK, Tutchko S, Egorin MJ. Phase I pharmacokinetic-pharmacodynamic study of 17-(allylamino)-17-demethoxygeldanamycin (17AAG, NSC 330507), a novel inhibitor of heat shock protein 90, in patients with refractory advanced cancers. Clin Cancer Res, 2005; 11(9):3385–91. CrossRef
Sequist L, Janne P, J S. In Inhibitor IPI-504, in Patients with relapsed and/or refractory stage iiib or stage iv non-small cell lung cancer (NSCLC) stratified by EGFR mutation status. AACR-NCI-EORTC International Conference on Molecular Targets and Cancer Therapeutics, 2007. CrossRef
Smith V, Sausville EA, Camalier RF, Fiebig HH, Burger AM. Comparison of 17-dimethylaminoethylamino-17-demethoxy-geldanamycin (17DMAG) and 17-allylamino-17-demethoxygeldanamycin (17AAG) in vitro: effects on Hsp90 and client proteins in melanoma models. Cancer Chemother Pharmacol, 2005; 56(2):126–37. CrossRef
Stebbins CE, Russo AA, Schneider C, Rosen N, Hartl FL, Pavletich NP. Crystal structure of an HSP90–geldanamycin complex: targeting of a protein chaperone by an antitumor agent. Cell, 1997; 89(2): 239-50. CrossRef
Solit DB, Osman I, Polsky D, Panageas KS, Daud A, Goydos JS, Teitcher J, Wolchok JD, Germino FJ, Krown SE, Coit D, Rosen N, Chapman PB. Phase II trial of 17-allylamino-17-demethoxygeldanamycin in patients with metastatic melanoma. Clin Cancer Res, 2008; 14(24):8302–7. CrossRef
Supko JG, Hickman RL, Grever MR, Malspeis L. Preclinical pharmacologic evaluation of geldanamycin as an antitumor agent. Cancer Chemother Pharmacol, 1995: 36(4):305–15. CrossRef
Sydor JR, Normant E, Pien CS, Porter JR, Ge J, Grenier L, Pak RH, Ali JA, Dembski MS, Hudak J, Patterson J, Penders C, Pink M, Read MA, Sang J, Woodward C, Zhang Y, Grayzel DS, Wright J, Barrett JA, Palombella VJ, Adams J, Tong JK. Development of 17-allylamino-17-demethoxygeldanamycin hydroquinone hydrochloride (IPI-504), an anti-cancer agent directed against Hsp90. Proc Natl Acad Sci USA, 2006; 103(46):17408–13. CrossRef
Taechowisan T, Puckdee W, Phutdhawong WS. Streptomyces zerumbet, a Novel species from Zingiber zerumbet (L.) Smith and isolation of its bioactive compounds. Adv Microbiol, 2019; 9(3):194–219. CrossRef
Taechowisan T, Samsawat T, Jaramornburapong C, Phutdhawong W, Phutdhawong WS. Antiviral activity of dopamine geldanamycin hybrids against influenza virus and association with molecular docking analysis. Int J Pharmacol, 2021; 17(1):1–14. CrossRef
Taechowisan T, Samsawat T, Puckdee W, Phutdhawong WS. Cytotoxicity activity of geldanamycin derivatives against various cancer cell lines. J Appl Pharm Sci, 2020: 10(6):12–21. CrossRef
Teo RD, Dong SS, Gross Z, Gray HB, Goddard WA. Computational predictions of corroles as a class of Hsp90 inhibitors. Mol Biosyst, 2015; 11(11):2907–14. CrossRef
Tian ZQ, Liu Y, Zhang D, Wang Z, Dong SD, Carreras CW, Zhou Y, Rastelli G, Santi DV, Myles DC. Synthesis and biological activities of novel 17-aminogeldanamycin derivatives. Bioorg Med Chem, 2004; 12(20):5317–29. CrossRef
Whitesell L, Mimnaugh EG, De Costa B, Myers CE, Neckers LM. Inhibition of heat shock protein HSP90-pp60v-src heteroprotein complex formation by benzoquinone ansamycins: essential role for stress proteins in oncogenic transformation. Proc Natl Acad Sci USA, 1994; 91(18):8324–8. CrossRef
Wrona IE, Gozman A, Taldone T, Chiosis G, Panek JS. Synthesis of reblastatin, autolytimycin, non-benzoquinone analogues: potent inhibitors of heat shock protein 90. J Org Chem, 2010; 75(9):2820–35. CrossRef