INTRODUCTION
The glycation process happens naturally in the human body as the process of aging. However, in diabetes mellitus (DM) patient, glycation process becomes uncontrollable, resulting in the accumulation of advanced glycation end products (AGEs) as firstly identified in 1986 (Fournet et al., 2018; Singh et al., 2014; Rahbar, 1968). The accumulation of AGEs and interaction of AGEs/receptor AGEs (RAGE) are the basis for the development of complications in DM, including cardiovascular disease, neurodegenerative disorder, chronic kidney disease, and ophthalmology disease (Sarwar and Gao, 2010). As in 2017, it was reported that 429 million patients suffered from DM with a financial burden of USD 727 billion, which estimated to grow larger in 2045, reaching 628.6 million patients (IDF, 2017).
The formation of AGEs occurs via three different parts, which are nonenzymatic Maillard reaction, polyol pathway, and lipid peroxidation (Nowotny et al., 2015). Then, AGEs will accumulate in various organs leading to the disruption of intracellular and extracellular protein structure and function (Singh et al., 2014). AGEs also initiate a signaling pathway mediated by several membrane receptors. The most studied AGEs receptor is the type 1 cell surface RAGE. The binding of AGEs and RAGE is responsible for reactive oxygen species generation and inflammatory responses through the expression of mitogen-activated protein kinase pathway, nuclear factor-kB, interleukin-6, tumor necrosis factor-α, intercellular adhesion molecule-1, and vascular cell adhesion molecule-2 (Grimm et al., 2018; Ott et al., 2014). Several studies of drugs that inhibit AGEs directly or indirectly by disrupting AGEs and RAGE have shown some promising results as the prevention of DM complication. Elparestat lowered the accumulation of AGEs effectively in peripheral neuropathy, whereas hesperidin lowered AGEs in the retina (Amore et al., 1997; Schmidt et al., 1994). The therapy using a synthetic inhibitor of AGEs, although promising, has the downside risk, which is a more prominent side effect. Therefore, the studies had leaned into searching for alternative traditional inhibitor for AGEs. Several studies on alternative traditional inhibitors, including grape seed proanthocyanidins extract (Cui et al., 2008), curcumin (Stefanska, 2012), Litsea japonica (Sohn et al., 2013), and Korea ginseng extract (Quan et al., 2013), had shown that these agents were comparable as AGEs inhibitors.
Centella asiatica (CA), a widely used traditional medicinal plant in Indonesia, was reported to have some clinical uses for preventing DM complications. Active compounds of CA consist of pentacyclic triterpenes, mainly asiatic acid, madasiatic acid, madecassic acid, asiaticoside, and madecassoside. Moreover, there were several other triterpenes, including betulinic acid, isothankunic acid, terminolic acid, and centellasapogenol A (Orhan, 2012). These active compounds have been proven to possess anti-inflammation and anticancer properties, which are beneficial for attenuating glycative process. Pentacyclic triterpenes also had anti-AGE and anti-RAGE agents based on several in vivo studies (Yin, 2015). Although the previous studies not explicitly focused on a specific active compound of CA, an in vitro study of CA proved its ability of lowering glycative activity. The CA purified extract had been shown to inhibit the Nε-carboxymethyl-lysine formation totally, which is one of the significant constituents of AGEs, induced by methylglyoxal (Maramaldi et al., 2014). This inhibition potential could be explored via in silico method to understand the mechanism of the action of CA better. The in silico method provides current development of the method in testing pharmacology hypothesis via measuring the affinity of a new molecule to a target and assessing the physiochemical properties (absorption, distribution, metabolism, and excretion) of new drugs (Ekins et al., 2007).
This study aims to explore the molecular docking of CA active compound as an inhibitor of AGEs and RAGE. Furthermore, molecular dynamic (MD) simulations were used in the study to assess the stability of binding complex. We also analyzed the probable molecular mechanisms of the action of CA active compounds on the AGEs and RAGE.
MATERIALS AND METHODS
This study was conducted using the software MarvinBean Suite®, Python Molecular Viewer® (PMV), PyRx-Autodock-Vina®, and Amber® 14. The software was operated in PC Intel® core i7 3770 3,5 GHz, Quad Core-8 threaded, RAM 32 GB DDR3 10600.
Ligand and protein preparation
The protein targets of AGEs and RAGE were generated from three-dimensional structure modeling in.pdb format from Protein Bank Data (PDB) (www.pdb.org) code with PDB ID: 3DKK for AGE and PDB ID: 4YBH for RAGE. The AGE protein structure (PDB ID 3DKK) named Aged Form of Human Butyrylcholinesterase inhibited by Tabun as the native ligand. The RAGE protein structure belongs to the crystal structure of the human RAGE ectodomain (VC1C2 fragment) in complex with human S100A6 as the native ligand. The three-dimensional structures were obtained from www.pdb.org. AGEs and RAGE underwent an optimization process using docking preparation, which consisted of hydrogen addition or current addition through Amber® 14 method. The residue preparation was facilitated according to default in Python Molecular Viewer® (PMV). The co-crystal ligand chain of AGEs and RAGE in the target was separated from the protein chain. The co-crystal ligand was selected as crystallography ligand according to the interaction model from PDB. The co-crystal ligands were prepared by hydrogen addition or charge addition. Nine of CA compounds were selected from PubChem® database search engine. The three-dimensional structure was then optimized by PMV®.
Docking validation
The prepared protein target and crystallography ligand were later put into PyRx-Autodock-Vina. Gridbox was set in protein receptor location with coordinate X:1,291 Y:7,504 Z:37,569 and dimension X:30,031 Y:28,790 Z:27,798.
Crystallography ligand proteins from PDB were optimized for the docking process. The optimization of proteins and ligands was conducted four times. After the docking process, ligands were saved and then measured against crystallography ligand. The measurement resulted in the root-mean-square deviation (RMSD). Ligands with RMSD < 2 were eligible for the next process. Proteins from docking process and derivatives from ligand were put into PyRx® program. The results were saved in.csv format and.sdf format. The analytical process was based on binding energy, binding affinity, and interaction residue from molecular docking. This study used molecular docking to investigate the possible mechanisms of interaction of CA compounds. The compound with a significant in silico score was deemed possible as an inhibitor.
Chimera program was used for optimization with Dunbrack Rotamer Library, which served for repairing incomplete protein residue. An improvement of protein residue charge was based on AMBER and also Chimera program. The purpose was to adjust the protein charge for biomolecular simulation.
Dynamic simulation
This study performed MD simulation using the Amber® 14 software package (Case et al., 2014). MD simulation was the prevailing sampling algorithms to investigate the interaction between a ligand and a protein target. The docked structure of AGEs (PDB ID: 3DKK) was reported as the fourth most active compound and novel chemicals with high activity among in silico studies. Therefore, we used these docked structures as the initial structures for MD simulations. The missing hydrogen atoms of the AGEs were added by the LEaP module of the Amber® 14 package to maintain the electroneutrality. Each complex was immersed into a cubic periodic box of TIP3P water model with at least 10ËšA distance around the complex (Wang et al., 2004). For the ligands, the GAFF parameter assignments were made by using Antechamber program, and then, the partial charges were assigned by using the AM1-BCC method (Hou et al., 2011; Klebanov et al., 1998). Afterward, four 10-ns results from MD simulations were carried out with the PMEMD program without limit in the isothermal, isobaric ensemble (NPT, P = 1 atm, and T = 310 K MD. The time step was set at 2 fs. The cutoff of 10ËšA was applied to treat nonbonding interactions.
RESULTS AND DISCUSSION
The compounds were screened based on the energy variation and the formation of the ligand–receptor structure, given by the binding constant and the Gibbs free energy (ΔG) values. We analyzed ligand–protein binding by evaluating their binding affinity, binding structure, binding energy, and residue component.
Binding energy values were calculated as the sum of intermolecular energies (kcal/mol), comprising hydrogen bond energy, Van Der Waals energy, desolvation energy, and electrostatic energy. Binding energy represents the energy released due to the interaction between ligand and receptor. The ligand with the least amount of binding energy had the best possibility to interact with protein target. The value of pKi represents the level of affinity of the compound to the molecular target.
This study used AGE target model from Protein Data Bank named Aged Form of Human Butyrylcholinesterase inhibited by Tabun with the PDB ID: 3DKK, which contained with 529 amino acid. This three-dimensional structure has resolution 2.31 Å, R-value work 0.197, and R-value observed 0.200. The active side or receptor site for this enzyme was formed from the incorporation of both molecule chains with the primary residue. This protein target’s binding site was located in Asn106, Asn188, Lys190, Se191, Arg219, Asn241, Asn245, Phe278, Pro281, Gln316, Asp324, Ser338, Arg341, Gly413, Tyr420, Arg465, Asn485, and Tyr500.
Based on the result of docking with AGEs (Table 1), asiaticoside bound to AGEs with the least binding energy -11.8253 kcal/mol and pKi 11.829. Asiaticoside bound to AGEs with 10 hydrogen bindings which involve binding site of Asp324, Asp375, Asp391, Tyr420, Tyr500, Lys513, Gln518, Tyr420, Tyr 500, and Gln518 (Table 2). Another significant compound binding is madasiatic acid with a binding energy -10.6724 kcal/mol. Madasiatic acid had hydrogen binding in site Asp324 and Tyr500. Madecassic acid bound with -10.1462 kcal/mol.
In visual observation through the docking simulation (Figure 1), the three observed ligands (asiaticoside, madasiatic acid, and madecassic acid) made similar conformations. Asiaticoside, madasiatic acid, and madecassic acid had similar conformations in the core chain and side chain, and the only differences are asiaticoside glucose chain and kiral compound. Brahmol showed the most distinctive conformation from other ligands due to Brahmol's low molecular weight and simpler side chain compared to the other compounds.
The top-ranked compounds with the lowest RMSD values were extracted to obtain reasonable conformation. Figure 2 shows the binding mode of native ligand, asiaticoside, and madasiatic acid with AGEs. It could be seen that the docking position of asiaticoside and madasiatic acid located in the same position and orientation as the native ligand to AGEs ligand-binding site. All these results indicated that although two of them were outside of the model AD, these compounds such as asiaticoside and madasiatic acid might have good performance to inhibit AGEs. The RMSF in Figure 3 showed that, in some amino acid residues for hydrogen bond, there is no significant fluctuation. The essential amino acid residues for hydrogen bond AGE asiaticoside are Asp324, Asp375, Asp391, Tyr420, Tyr500, Lys513, Gln518, Tyr420, Tyr 500, and Arg515. The essential amino acid residues for hydrogen bond AGE native ligand (L-peptide linking) are Asp324, Asp375, Asp391, Tyr420, Tyr500, Lys513, Gln518, Tyr420, Tyr 500, and Arg515. The essential amino acid residues for hydrogen bond AGE madasiatic acid are Asp375, Lys323, Val377, Tyr420, and Arg515.
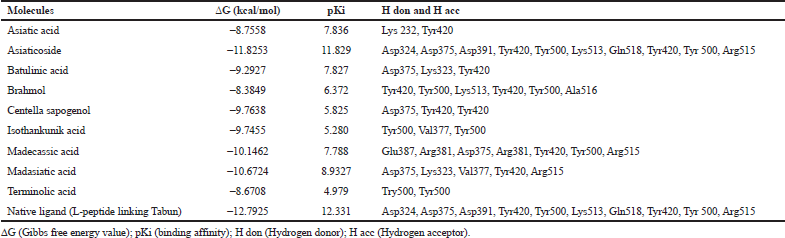 | Table 1: Binding energy and residue components of each ligand with AGEs. [Click here to view] |
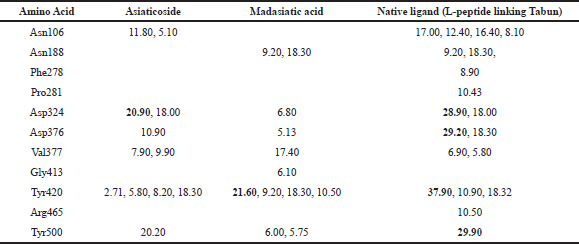 | Table 2: Occupation of hydrogen binding of complex AGEs [Click here to view] |
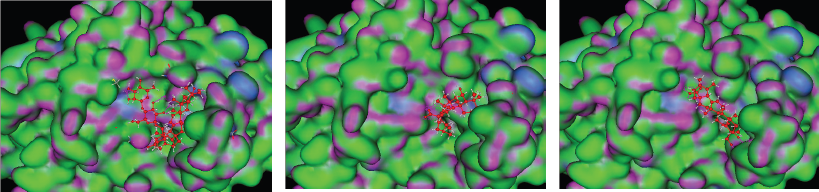 | Figure 1: 3D model binding of ligand (A) asiaticoside, (B) madasiatic acid, and (C) madecassic acid with AGEs. [Click here to view] |
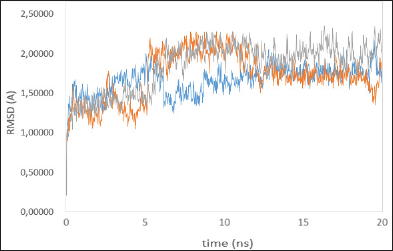 | Figure 2: Time series of the RMSD of the residues around 5ÌŠA of the ligand, native ligand (blue line), madasiatic acid (gray line), and asiaticoside (orange line). [Click here to view] |
To verify whether the studied systems reached equilibrium, the RMSD of all residues of the active site (residues within 5 ÌŠA around ligand) and the heavy atoms of the ligand from the initial structure were monitored to examine the dynamic stability of the systems and plotted against time, as shown in Figure 2. The three RMSDs had small fluctuations after 12 ns, implying that the studied systems had reached the stability. We used the last 20 ns to analyze the energy and binding modes for the four complexes.
Asn188, 278, 413, and 465 as a binding site which occupied Gly413 in madasiatic acid exhibited the inhibitory effect toward the protein activity. Nonetheless, MD simulation data annexed the inhibition level based on the number of occupancy binding to the active site (Asp324). Madasiatic acid, asiaticoside, and native ligand had occupied the active site Asp324 with a binding distance of 6.80, 20.90, and 28.90, respectively. Compared to native ligand, asiaticoside possessed a more stable inhibition level than madasiatic acid.
To obtain a detailed interaction between the two ligands and AGEs, we investigated the key residues during the binding process. The key residues were identified with the binding free energy decomposition calculated through the Molecular Mechanics/Generalized Born Surface Area (MM/GBSA) method. Energy decomposition was calculated from Van der Waals, electrostatic, solvation-free energy, and total energy contribution terms. The residues with energy contributions more than 1.5 kcal/mol were selected. The results are shown in Table 2. Asn106, Asp324, Asp376, Tyr420, and Tyr500 residues of AGEs made a significant contribution to the asiaticoside AGE complex. Meanwhile, Asn118 and Tyr500 residues made a significant contribution to the madasiatic acid AGE complex. The vast majority of the crucial residues of AGE was nonpolar, so it was reasonable to speculate that these residues were capable of forming more significant Van der Waals interactions with hydrophobic ligand. Therefore, the nonpolar attributes also exhibited a more favorable contribution of nonpolar interaction to the binding free energy.
Based on the result of docking with RAGE (Table 3), there were three ligands with the least binding energy for RAGE which are asiaticoside, isothankunik acid, and asiatic acid with a binding energy of -10.6125, -9.4469, and -9.1015 kcal/mol, respectively. The most common residues found from these interactions were Thr55, Arg57, and Glu103.
To explore the possible binding mode of the selected compounds, thus molecular docking was engaged to study the interaction in Figure 4 (especially, asiaticoside and isothankunik acid), together with the reported most active compound native ligand as a comparison, receptor of advanced glycation end products (RAGE) (PDB ID: 4YBH). The three-dimensional crystal structure of the human RAGE ectodomain (VC1C2 fragment) in complex with human S100A6 (PDB ID:4YBH) has a resolution of 2.40 Å with R-value work 0.195 and R-value observed 0.197. The Protein Data Bank has shown that RAGE was Aged Form of Human covered with 304 amino acid sequence length and no mutation. This target has binding site of Thr28, Thr55, arg57, Trp72, Arg78, Leu79, Asp93, Glu132, Lys169, His180, Asp201, His218, Arg228, and His270.
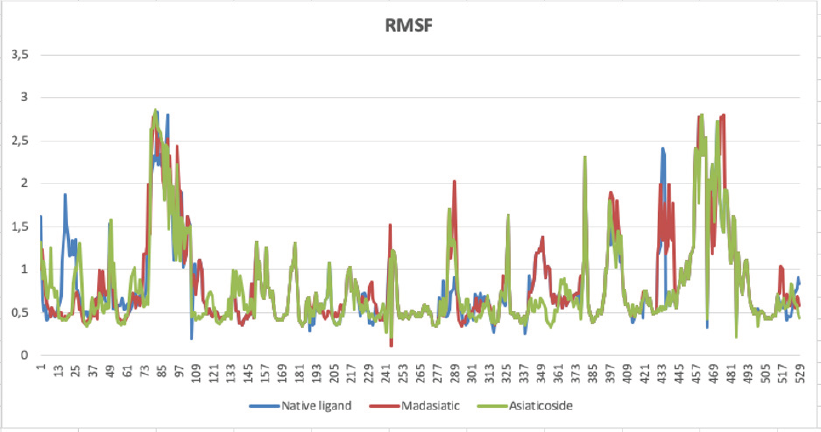 | Figure 3: Amino acid residues fluctuation (RMSF) of the native ligand (blue line), madasiatic acid (red line), and asiaticoside (green line). [Click here to view] |
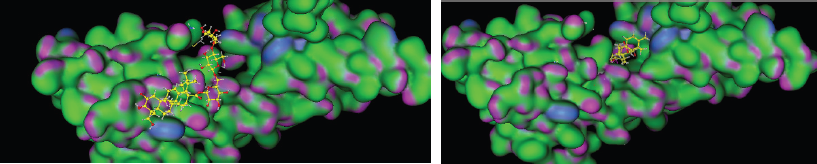 | Figure 4: 3D model binding of ligand (A) asiaticoside, and (B) ishothankunic acid with RAGE. [Click here to view] |
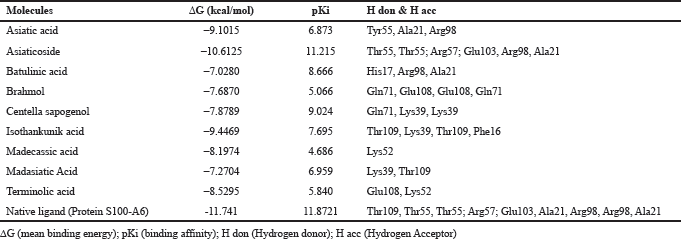 | Table 3: Binding energy and residue components of each ligand with RAGE. [Click here to view] |
Based on molecular dynamic evaluation (Table 2), we have discovered that asiaticoside has very close similarity in hydrogen bond occupancy with native ligand (L-peptide linking Tabun) in amino acid residues: Asp324, Val377, and Tyr500. Madasiatic acid has a strong similarity with the native ligand (L-peptide linking Tabun) in amino acid residues: Asn188 and Tyr420.
The interaction identified between CA and AGEs showed the CA’s role as AGE inhibitor. The individual substances tend to have different mechanisms as an AGEs inhibitor. Asiaticoside, madasiatic acid, and madecassic acid bound to the binding site of AGEs; thus, they are capable of preventing AGEs to bind to RAGE directly. The mechanism of modifying AGEs binding site to prevent an interaction with RAGE is similar to the mechanism of known AGEs inhibitor, carnosine, a dipeptide consisting of b-alanine and histidine. As noted in the carnosine study, the modification of the binding site also prevented cross-linking between AGE or AGE formation (Klebanov et al., 1998). In the previous study of CA compound, specifically studying asiaticoside, the result showed that asiaticoside lowered the level of AGEs (Schmidt et al., 1994).
In this study, we also discovered that asiaticoside, asiatic acid, and isothankunic acid bound to the binding site of RAGE. The mechanism of direct inhibition to the binding site of RAGE was previously studied in azeliragon. The capability of a ligand to bind to RAGE binding domain blocked the capability of another ligand to bind to RAGE (Burstein et al., 2014). In the study by Wang, the asiatic acid had lowered the level of RAGE in human skin keratinocyte (HaCat) (Wang, 2014).
Currently, the CA studies in animal model have provided evidence of CA role on oxidative stress. CA prevents cognitive deficit by the attenuation of oxidative stress and anti-inflammation activity, as well as restoring cholinergic function (Chiroma et al., 2019). Specifically, CA ability to enhance antioxidant activity, e.g., superoxide dismutase (Chiroma et al., 2019) catalase, and glutathione peroxidase (Ahmad Rather et al., 2019) while decreasing oxidative stress, e.g., malondialdehyde (MDA) (Chamaco-Alonso et al., 2019) and reactive nitrogen species (RNS) (Visnawathan et al., 2019) balances the antioxidant–pro-oxidant condition which is usually impaired in cognitive diseases, such as Alzheimer Disease. Centella asiatica was found to enhance antioxidant pathway through promoting nuclear transcription factor (erythroid-derived 2)-like 2 (NRF2) (Matthews et al., 2019) and activate Akt/GSK3β pathway (Ahmad Rather et al., 2019).
CONCLUSION
In this study, we applied the docking studies to CA active compounds with AGEs and RAGE. AGEs had been found to react with significant binding energy to asiaticoside, madasiatic acid, and madecassic acid with a binding energy of -11.8253, -10.6724, and -10.1462, respectively. Through MD simulation, asiaticoside and madasiatic acid bound into a similar site of native ligand with significant analyzed binding energy decomposition, thus confirming their role as an inhibitor of AGEs. CA active compound had also been proven to interact with AGEs and RAGE. The docking studies showed that the ligands RAGE bound to asiaticoside, isothankunik acid, and asiatic acid with a binding energy of -10.6125, -9.4469, and -9.1015 kcal/mol, respectively. Although this study proved the inhibition AGEs and RAGE by CA via in silico study, the findings were preliminary. These ligands can be further assessed for their effectiveness in clinical trials.
CONFLICT OF INTEREST
Authors declared that they do not have any conflicts of interest.
FINANCIAL SUPPORT
None.
REFERENCES
Ahmad Rather M, Justin-Thenmozhi A, Manivasagam T, Saravanababu C, Guillemin G, Essa M. Asiatic acid attenuated aluminum chloride-induced tau pathology, oxidative stress and apoptosis via AKT/GSK-3β signaling pathway in Wistar rats. Neurotox Res, 2019; 35(4):955–68. CrossRef
Amore A, Cirina P, Mitola S, Peruzzi L, Gianoglio B, Rabbone I, Sacchetti C, Cerutti F, Grillo C, Coppo R. Nonenzymatically glycated albumin (Amadori adducts) enhances nitric oxide synthase activity and gene expression in endothelial cells. Kidney Int, 1997; 51:27–35. CrossRef
Burstein AH, Grimes I, Galasko DR, Aisen PS, Sabbagh M, Mjalli AM. Effect of TTP488 in patients with mild to moderate Alzheimer’s disease. BMC Neurol, 2014; 14:12. CrossRef
Camacho-Alonso F, Torralba-Ruiz M, García-Carrillo N, Lacal-Luján J, Martínez-Díaz F, Sánchez-Siles M. Effects of topical applications of porcine acellular urinary bladder matrix and Centella asiatica extract on oral wound healing in a rat model. Clin Oral Investig, 2018; 23(5):2083–95. CrossRef
Case DA, Babin V, Berryman JT, Betz RM, Cai Q, Cerutti DS, et al.. Amber 14. University of California, California, 2014.
Chiroma S, Baharuldin M, Mat Taib C, Amom Z, Jagadeesan S, Ilham Adenan M et al. Protective effects of Centella asiatica on cognitive deficits induced by D-gal/AlCl3 via inhibition of oxidative stress and attenuation of acetylcholinesterase level. Toxics, 2019; 7(2):19. CrossRef
Cui XP, Li BY, Gao HQ, Wei N, Wang WL, Lu M. Effects of grape seed proanthocyanidin extracts on peripheral nerves in streptozocin-induced diabetic rats. J Nutr Sci Vitaminol (Tokyo), 2008; 54:321–8. CrossRef
Ekins S, Mestres J, Testa B. In silico pharmacology for drug discovery: applications to targets and beyond. British J Pharmacol, 2007; 152:21–37. CrossRef
Fournet M, Bonté F, Desmoulière A. Glycation damage: a possible hub for major pathophysiological disorders and aging. Aging Dis, 2018; 9(5):880. CrossRef
Grimm S, Ott C, Horlacher M, Weber D, Hohn A, Grune T. Advanced-glycation-end-product-induced formation of immunoproteasomes: Involvement of RAGE and Jak2/STAT1. Biochem J, 2012; 448:127–39. CrossRef
Hou T, Wang J, Li Y, Wang W. Assessing the performance of the MM/PBSA and MM/GBSA methods. 1. The accuracy of binding free energy calculations based on molecular dynamics simulations. J Chem Inf and Model, 2011; 51(1):69–82. CrossRef
International Diabetes Federation. IDF diabetes atlas. 8th edition, International Diabetes Federation, Brussels, Belgium, 2017.
Jakalian A, Bush B, Jack D, Bayly C. Fast, efficient generation of high-quality atomic charges. AM1-BCC model: I. Method J Comput Chem, 2000; 21(2):132. CrossRef
Klebanov GI, Teselkin YO, Babenkova IV, Rebrova OYU, Boldyrev AA, Boldyrev AA, Vladimirov Yu A. Effect of carnosine and its components on free-radical reactions. Membr Cell Biol, 1998; 12:89–99. CrossRef
Maramaldi G, Togni F, Franceschi E, Lati E. Anti-inflammaging and antiglycation activity of a novel botanical ingredient from African biodiversity (CentevitaTM). Clin Cosmet Investig Dermatol, 2014; 7:
1–9. CrossRef
Matthews DG, Caruso M, Murchison C, Zhu JY, Wright KM, Harris CJ, Gray NE, Quinn JF, Soumyanath A. Centella asiatica improves memory and promotes antioxidative signaling in 5XFAD mice. Antioxidants (Basel), 2019; 8(12):630. CrossRef
Nowotny K, Jung T, Höhn A, Weber D, Grune T. Advanced glycation end products and oxidative stress in type 2 diabetes mellitus. Biomolecules, 2015; 5(1):194–222. CrossRef
Ott C, Jacobs K, Haucke E, Navarrete-Santos A, Grune T, Simm A. Role of advanced glycation end products in cellular signaling. Redox Biol, 2014; 2:411–29. CrossRef
Orhan IE. Centella asiatica (L.) Urban: From traditional medicine to modern medicine with neuroprotective potential. Evid Based Complement Alternat Med, 2012; 2012:946259. CrossRef
Quan HY, Kim do Y, Chung SH. Korean red ginseng extract alleviates advanced glycation end product-mediated renal injury. J Ginseng Res, 2013; 37:187–93. CrossRef
Rahbar, S. An abnormal hemoglobin in red cells of diabetics. Clin Chim Acta, 1968; 22:296–98. CrossRef
Sarwar N, Gao P, Seshasai SR, Gobin R, Kaptoge S, Di Angelantonio E, Ingelsson E, Lawlor DA, Selvin E, Stampfer M, Stehouwer CD, Lewington S, Pennells L, Thompson A, Sattar N, White IR, Ray KK, Danesh J. Diabetes mellitus, fasting blood glucose concentration, and risk of vascular disease: a collaborative meta-analysis of 102 prospective studies. Lancet, 2010; 375:2215–22. CrossRef
Schmidt AM, Hasu M, Popov D, Zhang JH, Chen J, Yan SD, Brett J, Cao R, Kuwabara K, Costache G. Receptor for advanced glycation end products (AGEs) has a central role in vessel wall interactions and gene activation in response to circulating AGE proteins. Proc Natl Acad Sci USA, 1994; 91:8807–11. CrossRef
Singh VP, Bali A, Singh N, Jaggi AS. Advanced glycation end products and diabetic complications. Korean J Physiol Pharmacol, 2014; 18:1–14. CrossRef
Sohn E, Kim J, Kim CS, Lee YM, Jo K, Shin SD, Kim JH, Kim JS. The Extract of Litsea japonica reduced the development of diabetic nephropathy via the inhibition of advanced glycation end products accumulation in db/db mice. Evid Based Complement Alternat Med, 2013; 2013:769416. CrossRef
Stefanska B. Curcumin ameliorates hepatic fibrosis in type 2 diabetes mellitus - insights into its mechanisms of action. Br J Pharmacol, 2012; 166:2209–11. CrossRef
Viswanathan G, Dan V, Radhakrishnan N, Nair A, Rajendran Nair A, Baby S. Protection of mouse brain from paracetamol-induced stress by Centella asiatica methanol extract. J Ethnopharmacol, 2019; 236:474–83. CrossRef
Wang ZH. Anti-glycative effects of asiatic acid in human keratinocyte cells. Biomedicine, 2014; 4(3):19–28. CrossRef
Wang J, Wolf R, Caldwell J, Kollman P, et al. Development and testing of a general amber force field. J Comput Chem, 2004; 25(9):1157–74. CrossRef
Yin MC. Inhibitory effects and actions of pentacyclic triterpenes upon glycation. Biomedicine, 2015; 5(3):13. CrossRef