INTRODUCTION
Endometriosis has been known as the most frequent cause of pelvic pain in women during reproductive years. Endometriosis is a non-life threatening condition in which tissue that normally lines a woman’s uterus grows in other parts of the body, particularly on peritoneal tissues, bladder, ovaries, fallopian tubes, rectum, and other pelvic tissues. The prevalence of endometriosis is ~10%–15% among women of reproductive age (Crosignani et al., 2006; Mao and Anastasi, 2010; Vigano et al., 2004;) and up to 35%–50% among women with pelvic pain and/or infertility (Treloar et al., 2010). When endometriosis occurs, the eutopic endometrium experiences subtle abnormalities including biochemical reactions that increased the production of estrogen, prostaglandins, cytokines, and metalloproteinases (Bulun, 2009; Tjandrawinata and Rouli, 2017). These biological reactions resulted in pelvic pain, chronic pain, and fatigue which could lead to infertility.
In normal endometrium, progesterone exerts an antiestrogenic effect, in part by inducing 17β-hydroxysteroid dehydrogenase 2 (HSD17β2) which catalyzes the conversion of biologically potent estradiol to much less estrogenic estrone (Bulun et al., 2006). Progesterone acts on progesterone receptors (PRs) in endometrial stromal cells to increase the formation of retinoic acid as a paracrine factor, which induces HSD17β2 expression in endometrial epithelial cells. On the other hand, the estrogen level in the eutopic endometrium in women with endometriosis is higher than in normal endometrium, which may possibly be caused by the lack of interaction between progesterone and PR. As previously stated by Bulun et al. (2006), the increase of estrogen level in endometrial tissue is caused by progesterone resistance. Low level of progesterone is relatively related to prolonged release of estrogen and disordered proliferative endometrium. Prolonged estrogenic stimulation causes the endometrial glands to continue proliferating, becoming larger and more complex (Owings and Quick, 2014).
Selective PR modulator, such as asoprisnil, is a new class of PR ligands which have commonly been used in the studies of gynecological therapies such as uterine fibroids and endometriosis. This ligand can exert agonist and antagonist or mixed effects on various progesterone target tissues in vivo upon PR binding (Madauss et al., 2007). Asoprisnil leads to a less transcriptional activation than progestin and less transcriptional repression than antiprogestin. It also exhibits partial agonist/antagonist and tissue-selective effects in animals and humans. Although asoprisnil exhibits the potential to provide beneficial effects of progestins and antiprogestins, in this study, we focused on its agonist conformation as a model for molecular docking.
The development and use of the natural product have gained tremendous attention as a treatment of various disorders. Dexa Laboratories of Biomolecular Sciences (DLBS) has studied various herbals that exhibit potential effects on several diseases/disorders. For example is DLBS1425, a Phaleria macrocarpa extract which have antiproliferative and proapoptosis effects via eicosanoid pathway and downregulation of PI3K/AKT (Phosphatidylinositol-3-kinase/Protein kinase B) pathway (Tandrasasmita et al., 2010; Tjandrawinata et al., 2010). Other bioactive fraction from P. macrocarpa which called Proliverenol also has the potential ability in protecting cells from ethanol-induced hepatotoxicity (Berlian and Tandrasasmita, 2016). In vivo study by Tjandrawinata et al. (2015) presented that DLBS0533 which contains P. macrocarpa and Nigella sativa has an anti-inflammatory effect on mice model. An extract of pineapple (Ananas comosus) stem which named Tacorin and bioactive protein fraction isolated from Channa striata which called Striatin (DLBS0333) demonstrated acceleration of wound healing process by increasing cell proliferation (Rahayu et al., 2016; 2017). DLBS1442 is a proprietary and standardized semipolar bioactive fraction of P. macrocarpa (Scheff) Boerl fruit (Tjandrawinata et al., 2011). Previous clinical study indicated that DLBS1442 demonstrated its potential to treat symptoms of primary dysmenorrhea in premenstrual syndrome and alleviate endometriosis and/or adenomyosis related pain (Tjandrawinata et al., 2011, Wiweko et al., 2015). In vitro study by Tandrasasmita et al. (2015) reported that DLBS1442 significantly reduced the transcription level of the angiogenesis transcription factor, upregulated PR in RL95-2 cells, and induced cellular apoptosis in a dose-dependent manner. Therefore, DLBS1442 has been found to act as a potential agent to alleviate the symptoms of endometriosis via its antiangiogenic, anti-inflammatory, and proapoptotic activity. However, according to several activity studies of DLBS1442, the molecular mechanism of the active compounds that specifically play a role in generating its pharmacological effects has not yet been known.
In the present study, we investigated the bioactive compounds of DLBS1442 that may act as a PR agonist and evaluated its molecular activity. Our approach is to find the compound database of DLBS1442 and perform virtual screening. Database used in this study was obtained by the metabolomics approach. We hypothesized that the compound database of DLBS1442 could direct us to the compound that may possess an activity as a PR agonist.
MATERIALS AND METHODS
Preparation of receptor for virtual screening
X-ray crystal structure of the PR in complex with asoprisnil (PDB ID: 4A2J) was obtained from Protein Data Bank at www.rcsb.org (Lusher et al., 2012). X-ray crystal structure of the PR was separated from their bound ligand using Structure PrOtonation and REcognition System (SPORES) v1.3—mode splitpdb (Brink and Exner, 2009) and thus mol2 structures of 4A2J and asoprisnil were obtained. The crystal structure consists of two chains (A and B) where chain A was used. The binding site coordinates and the gridbox sizes were calculated based on binding site coordinates of asoprisnil in 4A2J using Protein-Ligand ANT Systems (PLANTS) v1.2—mode bind (Korb, 2009). The method also produced active site regions and active site amino acid residues as PLANTSactiveSite.mol2 and PLANTSactiveSiteResidues.mol2, respectively.
Comparing the binding pose of asoprisnil in X-ray crystal structure
Asoprisnil was re-docked to the crystal structure using PLANTS1.2_64bit—mode screen for 100 replicates. Cocrystal pose and re-dock pose of asoprisnil were superimposed in Pymol v2.10. Root mean square deviation (RMSD) of asoprisnil was calculated using command rms_cur. The visualization and RMSD calculation were performed in Pymol v2.10.
Decoy set and ligand set preparation
Decoy set (15,650) and ligand set (293) were obtained from DUD-E (dude.docking.org) in *.ism format (Mysinger et al., 2012). These compounds were converted into *.smi format then conditioned at pH 7.4. All files were converted into 3D (*.mol2) using Open-Babel v2.31 (O’Boyle et al., 2011). Decoys and ligands compound in *.mol2 were then processed using obconformer to get the best pose of each compound based on the Monte Carlo search. The *mol2 file was further processed using SPORES v1.3—mode reprot.
Virtual screening validation of 4A2J
Virtual screening was performed using PLANTS v1.2—mode screen with similar parameters used for redocking. For each compound, 50 binding poses were generated with triplicates replications. Other filtration was performed using Python-based Protein-Ligand Interaction Fingerprinting (PyPLIF) (Radifar et al., 2013), v0.1.1, which resulted in the Tanimoto Coefficient (TcPlif) score for each compound. Analysis result using Pylif v0.1.1 was then followed by Enrichment Factor (EF)1% rescoring based on the Chem Piecewise Linear Potential (ChemPLP) score from the best TcPlif value. The importance of hydrogen interaction of the ligand with GLN725 and ARG766 was also used as filtering system. Filtering was performed on compounds that exhibited hydrogen interaction with GLN725 and ARG766. EF1% was calculated based on the best ChemPLP score and TcPlif value.
Metabolomic study of DLBS1442
DLBS1442 was obtained from DLBS (Cikarang, West Java, Indonesia). Liquid chromatography–mass spectrometry/mass spectrometry (LC-MS/MS) analysis was performed using Agilent 1290 Infinity II LC with 6545 quadrupole-time of flight (QTOF) MS Detector. Separation was performed using XTerra MS C18 with 3.0 × 50 mm, 3.5 μm column. Acetonitrile (A) and ultrapure water (B) were used as the solvent. At a flow rate of 0.6 ml/minute, a gradient chromatographic system was performed using 2% solvent A into 100% solvent A for 11 minutes, followed by an equilibration of 2% solvent A over the next 4 minutes. MassHunter Workstation (B.06.01) was used as the processing software. The acquisition of MS/MS detector was performed at positive ion mode using Dual Agilent Jet Stream Electrospray ionization as an ion source.
LC-MS/MS result was processed using MZmine-2.32 (Pluskal et al., 2010), and the m/z of parent ion and fragmented ion was then collected. For compound prediction, we used METLIN at www.metlin.scripps.edu (Guijas et al., 2018). The simple search mode was used for early screening using the m/z value of the precursor ion. Predicted compounds from simple search were then continued with fragment similarity search using the m/z value of the precursor and fragment ions. Other references such as journals related to the metabolite compounds of DLBS1442 were also used in this metabolomics study.
Virtual screening of metabolomic study result
The structure of compounds from metabolomics study was obtained from PubChem (https://www.ncbi.nlm.nih.gov/pccompound) with *.sdf format, while the structure of new compound which was unavailable was generated using ChemDraw Ultra 12.0.2.1076 and saved as *.sdf format as well. Virtual screening was performed using PLANTS v1.2—mode screen using the validated protocol.
 | Figure 1. The superimposition between the docked conformation (yellow) and the cocrystal structure (turquoise blue) of the progesterone receptor-asoprisnil complex (4A2J). [Click here to view] |
RESULTS AND DISCUSSION
Virtual screening protocol validation
One commonly used method for validating docking program is pose selection whereby docking programs are used to re-dock into the target’s active site, compound with a known conformation and orientation, typically from co-crystal structure. In this study, the cocrystallized structure of PR in complex with asoprisnil (PDB ID: 4A2J) was used. The docking results showed that the binding conformation of the redocked asoprisnil reproduced the RMSD of less than 2.0 Ǻ, which was ranged from 0.962 to 0.973, as shown in Figure 1. Moreover, Figure 1 showed that the shifting of redocking results of asoprisnil (yellow) was insignificantly different compared to the cocrystal pose (turquoise blue). According to Hevener et al. (2009), programs that are able to obtain an RMSD value of lower than 2.0 Å were considered to exhibit a successful performance.
Virtual screening protocol validation for a PR agonist bind to asoprisnil crystalline structure was performed retrospectively. DUD/DUD-E is an example of a benchmark designed for validating virtual screening protocols and publicly available virtual screening test database (Mysinger et al., 2012). 15,650 decoys and 293 ligands in *.mol2 files from DUD-E were docked, and the protocol was then assessed for its ability to distinguish ligand from decoys. The assessed validation parameter was EF1% that is defined as the ratio of number of active compounds retrieved relative to the number of database molecules tested. Higher EF1% has been known to represent better protocol for ligand recognition because from the first 1% of the sorted database, the protocol was able to recognize ligands and exhibit higher ranks than decoys (Megantara et al., 2016).
Virtual screening protocol validation was performed using PLANTS (Korb, 2009). PLANTS has commonly been used for virtual screening as studied by Istyastono et al. (2015). All decoys and ligands were converted into smile format and prepared using Open Babel to adjust hydrogen atom into pH 7.4 that is similar to body’s condition. The smile format was then converted into three-dimension structure mol2 format using the same program, which could also be read by PLANTS. The mol2 format was processed using obconformer to get the best pose of each compound.
mol2 files of decoys and ligands were prepared using SPORES that performed structure recognition, and thus the three-dimension structures were adjusted to the algorithm suitable for PLANTS. The protonation by SPORES can either be done by adding the missing hydrogen atoms or as a complete reprotonation. List of amino acid residues which play a role in binding site was obtained using PLANTS. According to Lusher et al. (2012), oxosteroids, which include progesterone and asoprisnil, exhibit classic interactions to GLN725 and ARG766 with attachment to the hydrophobic pocket which consists of LEU715, LEU718, PHE794, LEU797, MET801, and TYR890.
Two scoring functions that were used for virtual screening validation consisted of ChemPLP and Tanimoto coefficient PLIF score (TcPLIF). PLANTS ChemPLP is an empirical scoring function that computes the fitness of protein−ligand binding by summing up the contributions of a number of individual terms, each represents an important energetic factor in protein−ligand binding (Liu et al., 2015). As a complementary method to ligand docking, TcPLIF can be applied to quantify the similarity of the predicted binding poses to a reference binding pose (Rácz et al., 2018).
Four protocols were used in the validation process. EF1% results from the protocols were tabulated in Table 1.
According to the results in Table 1, the best EF1% value, which is 7.16 was obtained on the protocol with hydrogen interaction with GLN725 and ARG766 residue and continued by rescoring based on ChemPLP score. This result was in accordance with the previous experiment by Williams and Sigler (1998) and Lusher et al. (2012) which found that hydrogen interaction with GLN725 and ARG766 is considered to be a vital interaction for both agonist and antagonist of PR functions.
Based on the best virtual screening protocol, the ChemPLP cutoff was −63.47. A compound could possibly be a ligand that acts as a PR agonist if it exhibits hydrogen interaction with GLN725 and ARG766 with the ChemPLP score −63.47.
Metabolomic study of DLBS1442
Metabolomic study was performed using LC-MS/MS with setting parameters as stated in the method. The gradient composition of the mobile phase consisted of 2% acetonitrile into 100% acetonitrile for 11 minutes, then continued with 4 minutes of 2% acetonitrile for equilibration. The low percentage of organic solvent at the beginning of analysis was intended to retain the DLBS1442 sample, so the compounds contained in DLBS1442 extract could be eluted gradually.
 | Table 1. EF1% value for each virtual screening protocol. [Click here to view] |
LC-MS/MS analysis provides information about precursor ions and fragment ions in positive ion mode [M+H]+. METLIN database was used to predict the structure of compounds from LC-MS/MS analysis results. Fragment similarity search in METLIN was used for compound identification approach.
Chromatogram of DLBS1442 showed that the highest peak gave a retention time of approximately 3.1 minutes. m/z value of the peak at retention time 3.04 minutes was 261.0762 and 423.1292. After mass fragmentation, precursor with m/z value of 423.1289 exhibited m/z value for fragmented ions of 261.0756, 167.0341, 121.0287, and 85.0286. However, identification in the METLIN database using both fragment similarity search and MS/MS spectrum match search did not gave any information about the m/z values. Literature study was also performed during the present metabolomics study. A study by Oshimi et al. (2008) stated that phalerin is one of the major compounds found in P. macrocarpa fruits and is believed to possess medicinal effects. Based on its structure, phalerin is a benzophenone glucoside which has been identified as 2,4’,6-trihydroxy-4-methoxybenzophenone-2-O-glucoside. After comparing MS data of DLBS1442 and phalerin standard, it was found that m/z value of 261.0761 and 423.1287 was also found in phalerin standard with similar retention time at 3.03 minutes (Fig. 2). Precursor ion with m/z value 423.1287 in phalerin was also fragmented into 261.0756, 167.0342, 121.0287, and 85.0286. This data suggests that precursor with m/z value 423.1289 at retention time 3.04 minutes in DLBS1442 might be phalerin.
Other predicted compounds from DLBS1442 were tabulated in Table 2. Due to the limitation of the database used, other references such as journals related to the metabolite compounds of DLBS1442 were also used for the metabolomics study. Ramdani et al. (2017) have identified several compounds in DLBS1442, which include glyceryl pentacosanoate, 1,7-dihydroxy-3,6-dimethoxyxanthone, 1,6,7-trihydroxy-3-methoxyxanthone, 2,4’,6-trihydroxy-4-methoxybenzophenone-2-O-glucoside, β-Sitosterol, Coumarin, and 2,3-dihydroxybenzoic acid.
Activity prediction of metabolomic study result as PR agonist
PR agonist activity from metabolomics study was predicted using virtual screening protocol which has previously been validated. Sample preparation, molecular docking parameters, and data processing were performed similar to the virtual screening protocol validation.
All identified compounds of DLBS1442 were used for activity prediction. Virtual screening result showed that there is only one compound that possesses ChemPLP ≤ −63.47 (Table 3). The lowest ChemPLP was obtained in glyceryl pentacosanoate with a ChemPLP score of −87.29. This compound is a small molecule that exhibits hydrogen interaction with GLN725 at 1.7 Ǻ and ARG766 at 2.0 Ǻ. These distances were closer compared to the asoprisnil 2.2 Ǻ and 2.4 Ǻ with GLN725 and ARG766, respectively (Fig. 3A and B). This closer distance for interaction with GLN725 and ARG766 could be supported with the low ChemPLP which indicates that glyceryl pentacosanoate could spontaneously interact with the PR. Figure 3C shows the best pose of glyceryl pentacosanoate as a PR agonist compared to asoprisnil.
 | Figure 2. Chromatogram of phalerin (1), MS (2), and MS/MS (3). [Click here to view] |
 | Table 2. Compounds identification from DLBS1442 based on metabolomic study. [Click here to view] |
 | Table 3. Virtual screening result from the metabolomic study of DLBS1442 compounds. [Click here to view] |
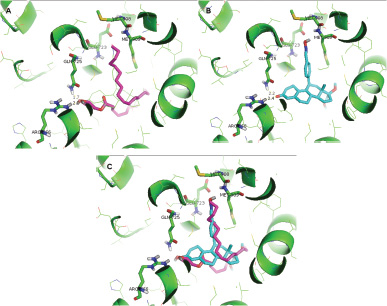 | Figure 3. Interaction of GLN725 and ARG766 with (A) glyceryl pentacosanoate and (B) asoprisnil. (C) Overlay position of glyceryl pentacosanoate and asoprisnil. [Click here to view] |
In silico study of DLBS1442 found that glyceryl pentacosanoate might have a role as a PR agonist. This result supported the previous study by Tandrasasmita et al. (2015) which found that DLBS1442 could also increase the expression of the PR. This finding shows the agonist-like activity of DLBS1442 to the PR, thus allowing better understanding of its molecular activity as anti-endometriosis in women. However, this finding requires further in vitro and in vivo approach to ensure the agonist activity of glyceryl pentacosanoate as the active compound of DLBS1442.
CONCLUSION
The virtual screening protocol for PR agonist has retrospectively been validated using PLANTS, PyPLIF, and DUD-E database. On the other hand, the metabolomics study of DLBS1442 also leads us to the specific compounds which might have a role for its activity. Virtual screening of the metabolomics study of DLBS1442 showed that only glyceryl pentacosanoate exhibits ChemPLP ≤ −63.47 and hydrogen interaction with GLN725 and ARG766. This indicates that glyceryl pentacosanoate could possibly play an important role as a PR agonist. Further approach by in vitro and in vivo studies would provide great support to prove its medical benefit for human life.
ACKNOWLEDGMENTS
The author would like to thank Recombinant Therapeutics Protein (RTP), Pharmacognosy and Extraction (PCE), Analytical (ANL) Sections of Dexa Laboratories of Biomolecular Sciences (DLBS), and staff member of Atma Jaya Catholic University for all technical supports. This work was in part supported by scholarship from DLBS.
CONFLICT OF INTEREST
Nil.
REFERENCES
Berlian G, Tandrasasmita OM, Tjandrawinata RR. Standardized bioactive fraction of Phaleria macrocarpa (Proliverenol) prevents ethanol-induced hepatotoxicity via down-regulation of NF-κB-TNFα-caspase-8 pathway. Asian Pac J Trop Biomed, 2016; 6(8):686–91. CrossRef
Brink T, Exner TE. Influence of protonation, tautomeric, and stereoisomeric states on protein−ligand docking. J Chem Inf Model, 2009; 49(6):1535–46. CrossRef
Bulun SE, Cheng YH, Yin P, Imir G, Utsunomiya H, Attar E, Innes J, Julie Kim J. Progesterone resistance in endometriosis: link to failure to metabolize estradiol. Mol Cell Endocrinol, 2006; 248(1-2):94–103. CrossRef
Bulun SE. Mechanism of disease endometriosis. N Engl J Med, 2009; 360:268–79. CrossRef
Crosignani PG, Olive D, Bergqvist A, Luciano A. Advances in the management of endometriosis: an update for clinicians. Hum Reprod Update, 2006; 12(2):179–89. CrossRef
Guijas C, Montenegro-Burke JR, Domingo-Almenara X, Palermo A, Warth B, Hermann G, Koellensperger G, Huan T, Uritboonthai W, Aisporna AE, Wolan DW, Spilker ME, Benton HP, Siuzdak G. METLIN: a technology platform for identifying knowns and unknowns. Anal Chem, 2018; 90(5):3156–64. CrossRef
Hevener KE, Zhao W, Ball DM, Babaoglu K, Qi J, White SW, Lee RE. Validation of molecular docking programs for virtual screening against dihydropteroate synthase. J Chem Inf Model, 2009; 49(2):444–60. CrossRef
Istyastono EP, Setyaningsih D. Structure-based virtual screening protocols to identify potent ligands for human adrenergic beta-2. Indonesian J Pharm, 2015; 26(1):20–8. CrossRef
Korb O. Efficient ant colony optimization algorithms for structure- and ligand-based drug design. Chem Cent J, 2009; 3(Suppl 1):O10. CrossRef
Liu J, Wang R. Classification of current scoring functions. J Chem Inf Model, 2015; 55(3):475–82. CrossRef
Lusher SJ, Raaijmakers HCA, Vu-pham D, Kazemier B, Bosch R, McGuire R, Azevedo R, Hamersma H, Dechering K, Oubrie A, van Duin M, de Vlieg J. X-ray structures of progesterone receptor ligand binding domain in its agonist state reveal differing mechanisms for mixed profiles of 11 β-substituted steroids. J Biol Chem, 2012; 287(24):20333–43. CrossRef
Madauss KP, Stewart EL, Williams SP. The evolution of progesterone receptor ligands. Med Res Rev, 2007; 27(3):374–400. CrossRef
Mao AJ, Anastasi JK. Diagnosis and management of endometriosis: the role of the advanced practice nurse in primary care. J Am Acad Nurse Pract, 2010; 22(2):109–16. CrossRef
Megantara S, Iwo MI, Levita J, Ibrahim S. Determination of ligand position in aspartic proteases by correlating tanimoto coefficient and binding affinity with root mean square deviation. J Appl Pharm Sci, 2016; 6(01):125–9. CrossRef
Mysinger MM, Carchia M, Irwin JJ, Shoichet BK. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J Med Chem, 2012; 55(14):6582–94. CrossRef
O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR. Open babel: an open chemical toolbox. J Cheminform, 2011; 1–14. CrossRef
Oshimi S, Zaima K, Matsuno Y, Hirasawa Y, Iizuka T, Studiawan H, Indrayanto G, Zaini NC, Morita H. Studies on the constituents from the fruits of Phaleria macrocarpa. J Nat Med, 2008; 62:207–10. CrossRef
Owings RA, Quick CM. Endometrial intraepithelial neoplasia. Arch Pathol Lab Med, 2014; 138:484–91. CrossRef
Pluskal T, Castillo S, Villar-Briones A, OrešiÄ M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics, 2010; 11:395. CrossRef
Rácz A, Bajusz D, Héberger K. Life beyond the Tanimoto coefficient: similarity measures for interaction fingerprints. J Cheminform, 2018; 10:48. CrossRef
Radifar M, Yuniarti N, Istyastono EP. PyPLIF: python-based protein-ligand interaction fingerprinting. Bioinformation, 2013; 9(6):325–28. CrossRef
Rahayu P, Agustina L, Tjandrawinata RR. Tacorin, an extract from Ananas comosus stem, stimulates wound healing by modulating the expression of tumor necrosis factor α, transforming growth factor β and matrix metalloproteinase 2. FEBS Open Bio, 2017; 7(7):1017–25. CrossRef
Rahayu P, Marcelline F, Sulistyaningrum E, Suhartono MT, Tjandrawinata RR. Potential effect of Striatin (DLBS0333), a bioactive protein fraction isolated from Channa striata for wound treatment. Asian Pac J Trop Biomed, 2016; 6(12):1001–7. CrossRef
Ramdani ED, Marlupi UD, Sinambela J, Tjandrawinata RR. Isolation and identification of compounds from Phaleria macrocarpa (Scheff.) Boerl fruit extract. Asian Pac J Trop Biomed, 2017; 7(4):300–5. CrossRef
Tandrasasmita OM, Lee JS, Baek SH, Tjandrawinata RR. Induction of cellular apoptosis in human breast cancer by DLBS1425, a Phaleria macrocarpa compound extract, via downregulation of PI3-kinase/AKT pathway. Cancer Biol Ther, 2010; 10(8):814–24. CrossRef
Tandrasasmita OM, Sutanto AM, Arifin PF, Tjandrawinata RR. Anti-inflammatory, antiangiogenic, and apoptosis-inducing activity of DLBS1442, a bioactive fraction of Phaleria macrocarpa, in a RL95-2 cell line as a molecular model of endometriosis. Int J Womens Health, 2015; 7:161–9. CrossRef
Tjandrawinata RR, Arifin PF, Tandrasasmita OM, Rahmi D, Aripin A. DLBS1425, a Phaleria macrocarpa (Scheff.) Boerl. extract confers anti proliferative and proapoptosis effects via eicosanoid pathway. J Exp Ther Oncol, 2010; 8(3):187–201.
Tjandrawinata RR, Djunarko I, Fenty, Hendra P. Anti-inflammation effects of bioactive fraction DLBS0533 containing Phaleria macrocarpa and Nigella sativa on animal model. Int J Pharm Pharm Sci, 2015; 7(1):408–11.
Tjandrawinata RR, Nofiarny D, Susanto LW, Hendri P, Clarissa A. Symptomatic treatment of premenstrual syndrome and/or primary dysmenorrhea with DLBS1442, a bioactive extract of Phaleria macrocarpa. Int J Gen Med, 2011; 4:465–76. CrossRef
Tjandrawinata RR, Rouli HC. A role for Phaleria macrocarpa (Scheff) Boerl. Extracts in the management of women’s pathological conditions: a research review. Int J Pharm Pharm Sci, 2017; 9(3):7–12. CrossRef
Treloar SA, Bell TA, Nagle CM, Purdie DM, Green AC. Early menstrual characteristics associated with subsequent diagnosis of endometriosis. Am J Obstet Gynecol, 2010; 202(6):534.e1-6. CrossRef
Vigano P, Parazzini F, Somigliana E, Vercellini P. Endometriosis : epidemiology and aetiological factors. Best Pract Res Clin Obstet Gynaecol, 2004; 18(2):177–200. Wiweko B, Puspita CG, Tjandrawinata R, Situmorang H, Harzif AK, Pratama G, Sumapriya K, Natadisastra M, Hestiantoro A. The effectiveness of Phalleria macrocarpa bioactive fraction in alleviating endometriosis and/or adenomyosis related pain. eJKI, 2015; 3(1):51–6. CrossRef
Williams SP, Sigler PB. Atomic structure of progesterone complexed with its receptor. Nature, 1998; 393:392–5. CrossRef