INTRODUCTION
Salmonella typhimurium is a bacterium classified as a Gram-negative, facultative anaerobe that is associated with the Salmonella subspecies I Serogroup D and has several genotypic and phenotypic variants with the Salmonella genus. Most of the traits of the Salmonella genus are shared by S. typhimurium [1]. To date, about 2,500 distinct serotypes or serovars have been found within two species, Salmonella enterica and Salmonella bongori. Salmonella is a common and robust bacterium that may persist in dry conditions and months in water. Every year, about 1 out of every 10 individuals, becomes ill, resulting in the loss of 33 million healthy lives per year [2]. In the year 2000, it was projected that over 21.6 lakh cases of typhoid disease were recorded globally, resulting in 2.16 lakh deaths, with Asia representing more than 90% of the incidence and death [3]. In 2010, the incidence of enteric fever in South Asia was 394.2 incidents per 1 lakh candidates. Salmonella enterica serovars S. typhimurium (typhimurium) or S. paratyphi (Paratyphi) A, B, or C cause enteric fever, a severe systemic infectious disease caused by human-restricted pathogens. The disease is transmitted through water, food, and direct person-to-person contact [4]. A prolonged fever has been observed in Salmonella infections, with a temperature of 39°C–40°C (103°F–104°F), with stomach ache, diarrhea, cough, constipation, headache, and loss of appetite [5]. Salmonella typhimurium is transmitted by touching your mouth before washing your hands when using an infected washroom, having to eat seafood from a source of water polluted by infected fecal matter or urinating, having to eat raw vegetables fertilized with human feces, engaging in oral and anal intercourse with a partner who is a carrier of S. typhimurium bacteria [6]. Salmonella microbes colonize the small and large intestines after passing through the stomach after being consumed. The bacteria subsequently infect and proliferate within the mucous membrane of the gut. The bacteria have the capability to infiltrate the lymphoid tissues of the gastrointestinal tract and migrate into the bloodstream. Transmission via the bloodstream is rare, happening in less than 5% of infections, and is dependent on host factors and the pathogenicity of the Salmonella strain. Any organ can become infected if the bacteria spread to the bloodstream (e.g., liver, gallbladder, and bones) [5]. In the northeast region, typhoid fever is a very common disease transmitted from contaminated food and infected water. The traditional culture-based methods for the exposure of Salmonella are time-consuming and may not always be reliable [7]. Recently, molecular techniques such as polymerase chain reaction (PCR) have been developed and used as alternative or supplementary methods for the detection of Salmonella. PCR-based methods have been shown to be highly sensitive and specific for the detection of Salmonella in various types of clinical samples, such as blood, stool, and environmental samples [8].
If the affected individual has a compromised immune system, they are more likely to experience high fevers or bloody stools, especially if they are a baby, young child, older adult, or someone else, which appears to be leading to dehydration, as evidenced by symptoms such as dry mouth and tongue, peeing less frequently than normal, and dark urine [9]. Infection with Salmonella is apparent in different places and the body has several built-in defenses against Salmonella infection in cases of stomach or intestinal issues. Strong stomach acid, for instance, can eradicate many Salmonella bacterial strains [10]. The patient might get dehydrated after a Salmonella infection if he/she does not drink enough to replenish the fluids you lose via diarrhea [5]. Bacteremia, which occurs when Salmonella infection reaches the circulation, can infect tissues all throughout the body. Reactive arthritis can also cause joint pain, eye discomfort, and painful urination [10].
PCR stands as a laboratory method engineered to generate numerous copies of a precise DNA segment, thereby facilitating meticulous examination. In this method, short synthetic DNA fragments, referred to as primers, are utilized to identify and amplify a particular segment of the genome. Following primer targeting, successive cycles of DNA synthesis are executed to amplify the targeted segments [11]. These sources describe various methods of detecting Salmonella using PCR techniques. The studies include the detection of Salmonella in blood [12], in environmental samples [13], and in raw milk [14].
A few years ago there were no accurate, fast, or sensitive procedures for clinical identification of disease-causing Salmonella species and these methods were used by various researchers. The disadvantages of previous methods for the disclosure of Salmonella species are not fully accurate and time-saving methods. For example, isolation of the causal organism remains the most successful diagnostic strategy in suspected typhoid fever, which decreases the progressing sickness, and blood has been the predominant sample for Salmonella serovars typhimurium culture since the eighteenth century [15,16]. Blood culture sensitivity peaks during the initial week of infection and decreases as the disease progresses and the blood culture method is prolonged and requires not less than 1 week before the organism can be identified [17–19]. Various factors such as insufficient laboratory medium, inherent bactericidal properties of blood, the quantity of blood obtained for culturing, antibiotic presence, and the period of blood collection can affect the challenge of isolating the causative agent from blood specimens [19].
PCR TYPES AND THEIR SIGNIFICANCE
The sensitive and precise identification of several diseases, including Salmonella, has been made possible by PCR, which has completely transformed the area of molecular diagnostics. Mullis [20] invented the molecular biology method known as PCR, for which he got the Nobel Prize in Chemistry in the year of 1993 . This technology, which amplifies certain DNA sequences, has the potential to identify extremely small amounts of genetic material. The DNA polymerase used in this method repeats the target DNA on a regular basis and rapidly increases the number of copies [11]. In the context of Salmonella detection, PCR offers several advantages. It allows for the quick and highly sensitive identification of Salmonella DNA in diverse samples, ranging from blood specimens [21] to environmental samples [22]. The specificity of PCR is attributed to the use of primers designed to match unique regions of the Salmonella genome [21]. This specificity is crucial for distinguishing between different Salmonella serovars, such as typhimurium and paratyphi [23]. Different types of PCR and their various applications in the field of biological sciences are discussed below.
Reverse transcription-polymerase chain reaction (RT-PCR)
RT-PCR stands as a pivotal molecular diagnostic tool for detecting RNA-based infections, exemplified in Salmonella identification. Comprising reverse transcription converting RNA to complementary DNA and PCR amplification for specific RNA sequence detection, real-time monitoring enhances precision [24]. Its specificity facilitates precise strain differentiation [23]. RT-PCR is very sensitive, identifying low Salmonella RNA levels even in complex materials such as blood, providing quick insights into infection severity [12]. Despite its advantages, RT-PCR poses challenges and complexity leading to potential errors, contamination risk, cost factors, and susceptibility to inhibitors in complex samples pose limitations [22]. Nonetheless, RT-PCR’s versatility extends across varied sample types, affirming its applicability and utility in diverse scenarios [20,25]. In the final analysis, RT-PCR appears as a powerful technology for understanding and controlling Salmonella infections because of its sensitivity, specificity, and real-time capabilities.
Multiplex polymerase chain reaction (Multiplex PCR)
Multiplex PCR serves as a sophisticated molecular biology technique designed to amplify multiple target DNA sequences simultaneously, facilitating the comprehensive detection of distinct Salmonella serovars in a single reaction [24,26]. Multiplex PCR, which uses distinct primer sets for different Salmonella DNA sections, enables the simultaneous amplification of several serovars, providing a thorough assessment of pathogen presence in a sample [24]. Notably, its ability to simplify diagnostics by combining many responses into a single test saves time and costs [26]. The reactivity and accuracy of multiplex PCR allow for the exact separation of Salmonella strains within complicated samples [12]. Despite its benefits, careful tuning is essential to balance target amplification, taking into account possible difficulties such as primer reactions and competition [24]. Multiplex PCR, which is extensively used across a wide range of sample types, including blood and environmental samples, is a versatile and effective approach for researching Salmonella serovars in a variety of contexts, adding to a better knowledge of the pathogen’s prevalence.
Nested polymerase chain reaction (Nested PCR)
Nested PCR is a sophisticated molecular biology technique employing two consecutive PCR processes, and is widely utilized in Salmonella detection to enhance sensitivity and specificity in DNA amplification [21,23]. This method is especially successful for detecting small levels of Salmonella DNA, which is critical in samples with few DNA templates [23]. Despite its sensitivity benefits, Nested PCR presents challenges, including a higher risk of contamination due to the two-step amplification process, necessitating strict laboratory protocols [23]. On the other hand, careful primer design and optimization add complexity, and considerations of potential cross-contamination, and increased resource needs are essential [21]. Widely applicable, Nested PCR is applied in diverse samples to showcase its adaptability and make it a valuable technique in scenarios requiring the identification of minimal Salmonella.
Real-time polymerase chain reaction (qPCR)
QPCR emerges as a sophisticated technology in molecular biology with significant advantages for Salmonella detection and monitoring in diverse samples [24,12]. Operating by monitoring DNA amplification in real-time through fluorescent dyes or probes, it provides precise, quick, and quantitative insights into the presence and severity of Salmonella infections [24]. Notably, it exhibits high sensitivity, enabling the identification of low amounts of Salmonella DNA in various materials, such as blood and environmental samples [12]. However, the approach has difficulties, including cost constraints due to specialized equipment and reagents, which restrict accessibility in resource-limited contexts [22] and this PCR requires skilled staff for correct execution and interpretation of results [23]. Despite drawbacks, real-time PCR’s adaptability and reliability in identifying Salmonella across diverse samples underscore its significance in comprehending and managing Salmonella infections, leveraging sensitivity, real-time monitoring, and quantification.
Conventional polymerase chain reaction (Conventional PCR)
Conventional PCR stands as a foundational molecular biology technique for the amplification of specific DNA sequences, playing a crucial role in Salmonella detection [24,12]. In addition, conventional PCR’s adaptability allows for the identification of specific Salmonella serovars [12]. Despite its advantages, conventional PCR has some limitations, its sensitivity is lower as compared to other modern PCR methods, which potentially affects the detection of Salmonella [12]. Contamination issues with this approach need stringent laboratory protocols. Its capacity to detect infection severity is hampered by the lack of real-time monitoring and quantification of amplified DNA [24].
DIFFERENT APPLICATION OF PCR FOR Salmonella DETECTION
In this study, we assessed the current literature on the utilization of PCR for the identification of Salmonella in blood samples. These studies have to be reviewed to use a variety of PCR-based methods, including multiplex PCR, RT-PCR, and nested PCR to detect Salmonella species, with a focus on S. typhimurium and paratyphi A. Considering the difficulties in diagnosing typhoid fever using blood culture and serological approaches, PCR techniques have lately been used. Since the initial study, in the year 1993, the PCR as an investigative tool for typhoid fever, when Song et al. [27] effectively amplified the flagellin gene (fliC-d) of Salmonella serovars typhimurium in altogether cases of culture proved typhoid fever and then other studies have been published by various researchers [28,29].
The genome of an organism is encoded in DNA molecules, but analyzing it necessitates an enormous amount of DNA. In 1985, Mullis [20] discovered that a small quantity of DNA could be duplicated in vast numbers over a short period of time using PCR. The basic principle of PCR is to heat and separate DNA’s two strands and also bind the DNA-building components supplied to each strand. The enzyme DNA polymerase is used to create new DNA chains, and the method is able to be reused. Forensic science and medical research have both used PCR extensively [20].
Some studies focus on the recognition of definite serovars of Salmonella, such as S. typhimurium [8,19,30] and S. paratyphi A [31] by using PCR techniques. Other studies also described the uses of multiplex PCR for concurrent identification of different Salmonella species [32,33] as well as diagnosis of typhoid pathogens [26]. Different researchers also revealed the application of RT-PCR for the quick recognition of various Salmonella species [13,33].
A DNA sequence serves as a genetic marker located at a known position on a chromosome that is employed for tracking the transmittal of a specific trait or disease in a family. By analyzing the transmission of the genetic marker through generations of a family, scientists can infer the location of the gene responsible for the trait or disease in question. Genetic markers are particularly useful when studying complex diseases, where multiple genes may contribute to the condition. In these cases, identifying the specific genes involved can be challenging, but genetic markers can help narrow down the search [23].
The genetic markers used for detecting Salmonella and various studies included genes such as IroB [12], flagellin [28], and others which are shown in Table 1. To detect the Salmonella organism from a blood sample, Ganesan et al. [12] utilized the iroB gene, which is responsible for the biosynthesis of salmochelins-iron-chelating compounds produced by Salmonella to acquire iron from the host. Khokhar et al. [8] identified highly conserved markers, including STY0307, intragenic region SSPA1732a-SSPA1724, STY0962, and STY2513, that indicate the high-risk lineages of S. typhimurium and paratyphi A. Zhou and Pollard [19] employed the fliC-d gene, which codes for flagellin and is involved in Salmonella motility. Kasturi and Drgon[13] used multiple genes, including invA, Salmonella-differentiating fragment 1 (Sdf-1), prt, tyv, and IAC, to identify Salmonella species. In addition, Zhou et al. [31] utilized the fliC-a gene specific to Salmonella paratyphi A. Park and Ricke[32] employed genes STM3098, STM4057, and STM4497 specific to Salmonella genus, Salmonella subsp. I, S. typhimurium, as well as the SEN0997 gene specific to Salmonella enteritidis and the restriction enzyme (ACF69659), respectively, to distinguish between different serovars of Salmonella. Their findings also demonstrated that multiplex PCR and qPCR may deliver speedy and reliable results for detecting and quantifying Salmonella in a variety of samples.
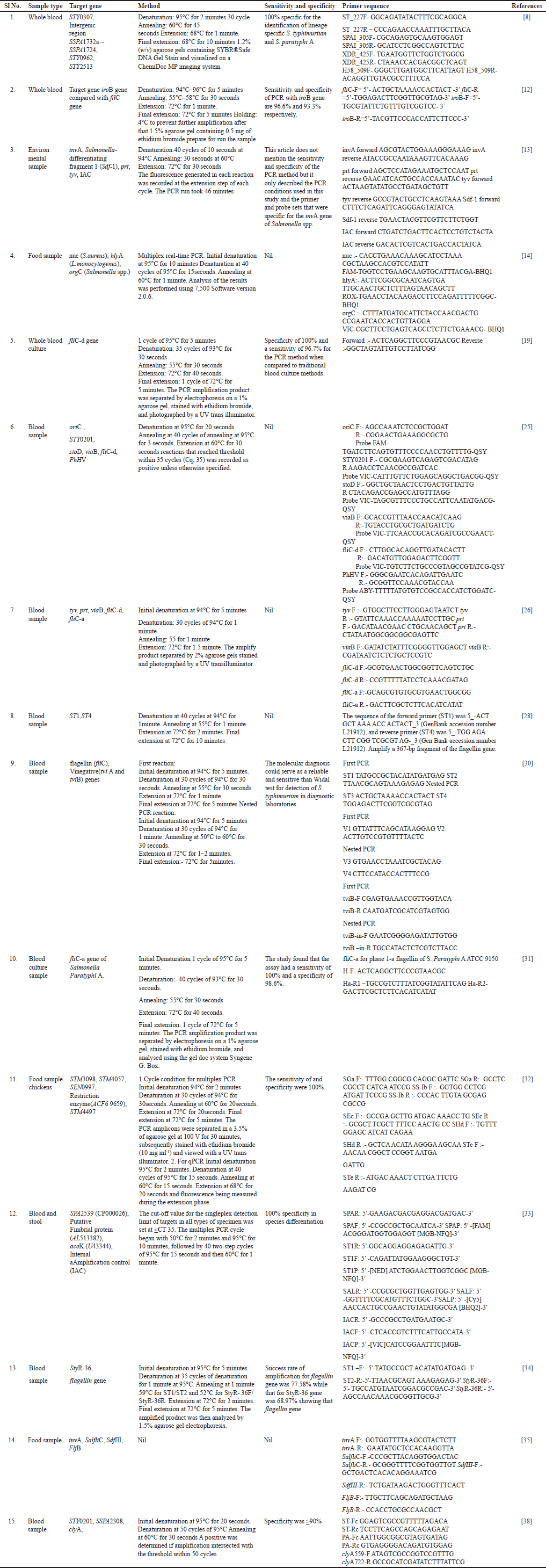 | Table 1. Salmonella species isolated from different sources and their target gene with methodology, sensitivity, primers and references. [Click here to view] |
The importance of gene markers in distinguishing salmonella species
Different gene markers serve a crucial function in the precise detection and isolation of Salmonella species, providing vital insights into their taxonomy and pathogenicity (Table 1). The iroB gene is required for bacterial siderophore production, which is required for iron uptake, and so provides insight into Salmonella iron metabolism and pathogenicity [12]. The filC gene contributes to flagellar biosynthesis, influencing Salmonella motility and pathogenicity and helping in the differentiation of motile and nonmotile forms [8]. The STY0307, SSPA 1732a-SSA 1724, STY0962, and STY2513 genes are unique to S. typhimurium and serve as markers for specific identification, allowing for quick and precise detection of this pathogenic strain [19,32]. Furthermore, the Sdf-1 gene acts as a significant marker for Salmonella species identification, assisting in molecular tests for their differentiation [12]. Prt, Tyv, STM3098, STM4057, SEN0997, and SPA2539 genes contribute to a complete knowledge of disease transmission by Salmonella [21,13]. The ST1, SAL, StyR-36, tyv, prt, viaB, and tviA genes are essential for diagnosing typhoid fever and understanding the genetic basis of S. typhimurium infections [34]. Toxin-producing and virulence genes such as nuc, hlyA, orgC, SalfliC, SdfIII, FljB, and oriC provide essential insights for assessing Salmonella’s pathogenic potential and developing targeted therapeutics [30,35].
Advantages and disadvantages of PCR methodology
It is notable for its capacity to identify large quantities of DNA amplified from a single or a few baseline sequences. Conventional PCR generates qualitative outcomes as opposed to quantitative ones. The benefits of PCR testing are that it is a chemical procedure that swiftly and exponentially increases the target nucleic acid. This method can generate thousands or even millions of replicates of a specific piece of RNA or DNA. This allows researchers and organizations to take only a small quantity of genetic material and amplify it to a sufficient volume for sequencing, analysis, or evaluation. By PCR technique we get faster results, shorter time to optimal therapy, improve treatment decisions, avoid unnecessary antibiotics, support antimicrobial stewardship efforts, reduce unnecessary testing, and reduce healthcare costs [22].
Previous sequencing data is required to overcome the limitations of PCR when creating primers. Therefore, the use of PCR is limited to identifying the presence or absence of a recognized pathogen or gene. Another observation is that PCR primers may bind to DNA sequences that are similar, though not perfectly identical to the target DNA. In addition, the DNA polymerase can incorporate erroneous nucleotides into the PCR sequence, but only in a very small amount [24]. The detection of Salmonella in various samples, such as food, blood, and environmental samples, has been greatly aided by PCR-based methods in recent years.
One of the frequently employed PCR-based procedures for the identification of Salmonella is the use of specific gene targets, such as the iroB gene for the exposure of Salmonella in blood specimens [12]. Other research findings have developed multiplex PCR assays that can detect multiple lineages of Salmonella, such as high-risk lineages of S. paratyphi A and S. typhimurium [8] as shown in Table 1.
Several studies have employed PCR-based methods to detect Salmonella in blood culture samples. Zhou and Pollard [19] emphasized that the PCR system is not only rapid but also highly responsive to identifying S. enterica serovars typhimurium from blood culture samples. In addition, a PCR assay designed for blood culture samples had been developed to identify S. paratyphi from clinical samples [31].
In addition to blood samples, PCR-based methods have been used for the detection of Salmonella from other forms of specimens. Kasturi and Drgon [13] developed a RT- PCR technique for the detection of Salmonella from the environmental samples. Park and Ricke [32] created a multiplex PCR test for the immediate identification of Salmonella sub species I, Salmonella genus, Salmonella Heidelberg, S. enteritidis, and S. Typhimurium because these Salmonella serovars were the most often isolated from poultry products. Some of the studies have also developed PCR-based methods for the recognition of definite serovars of Salmonella. In recent years, there has been a development of PCR-based methods for detecting Salmonella in different clinical samples, including blood. These methods have been shown to be exceedingly sensitive and exact, making them valuable utensils for the rapid diagnosis of Salmonellosis. One of the studies by Ganesan et al. [12] utilized PCR to detect Salmonella in blood samples using the iroB gene as a targeted gene and it was found that their process had a sensitivity of 100% and a specificity of 99.7%, making it a highly accurate method for the identification of Salmonella from blood sample. A multiplex PCR assay was created by Khokhar et al. [8] to identify lineages of S. typhimurium and Paratyphi A with high risk. The sensitivity of this assay was 97.5% and the specificity was 100% for these pathogens. The authors also found that their assay was able to detect these pathogens in blood samples from patients with established typhoid fever. Zhou and Pollard [19] developed a blood culture PCR technique for the uncovering of S. enterica serovar typhimurium. They discovered that their approach had a sensitivity of 96.7% and a specificity of 99.4%, and that was able to detect the pathogen from blood samples from patients with confirmed typhoid fever. Kasturi and Drgon [13] developed an RT-PCR method for the exposure of Salmonella from environmental samples. They found that their process had a sensitivity of 100% and a specificity of 99.8%, making it a highly accurate method for the recognition of Salmonella from these types of available samples. In addition to the findings of Salmonella from blood, several studies have also focused on the detection of specific serovars of Salmonella, such as S. typhimurium and Paratyphi A.
Overall, the present findings have shown that PCR-based methods are highly sensitive and specific for the detection of Salmonella from blood specimens. Numerous research findings have reported detection limits as low as a single Salmonella cell per reaction making PCR a highly sensitive method for detecting this pathogen in blood samples. In addition, several studies have reported high levels of specificity for PCR-based methods, with no false positives being reported in the studies reviewed. The use of multiplex PCR has been proven effective for the concurrent detection of multiple Salmonella serovars. Studies by Park and Ricke [32] and Ali et al. [26] have demonstrated the effectiveness of multiplex PCR for the immediate detection of Salmonella genus, S. enteritidis, Salmonella subspecies I, S. typhimurium in food and environmental samples. Similarly, Khokhar et al. [8] have developed a multiplex PCR test for the recognition of high-risk lineages of S. typhimurium and paratyphi A in human blood samples. The RT-PCR has also been shown and reported to be an active technique for the recognition of Salmonella from blood specimens. Blood culture is presently the gold standard for typhoid fever treatment; however, it is time consuming and takes several days to isolate and identify pathogenic organisms. In response to this, Zhou and Pollard [19] created a rapid and most accurate blood culture PCR technique for detecting S. typhimurium, enabling same-day therapy following an appropriate diagnosis of typhoid.
RT-PCR is chosen over standard PCR procedures because it enables real-time monitoring of target DNA amplification, which speeds up and simplifies the process [36,8]. RT-PCR applications include clinical microbiology, virology, and food microbiology, and countless tests for the detection and quantification of numerous pathogenic microorganisms have been reported by employing this technique [37,36]. The multiplex method has the advantage of reducing pipetting mistakes and allowing both the DNA and RNA to be amplified at the same time under the same conditions, in comparison to monoplex techniques, the duplex test yields more precise and accurate data [36]. Multiplex PCR is also used for Salmonella detection, which enables concurrent detection of numerous targets in a solitary reaction [8]. This method is useful for detecting different Salmonella serovars and differentiating Salmonella from other bacteria.
CONCLUSION
In conclusion, the detection of Salmonella from blood samples by applying PCR techniques is a highly accurate and specific method for the diagnosis of typhoid fever and other salmonellosis. This review demonstrates the utility of various PCR assays, such as those using the iroB gene, multiplex PCR, and RT-PCR for the detection of Salmonella from blood specimens. In addition, the use of specific molecular markers, such as the flagellin gene, can aid in the accurate identification of Salmonella serovars. Overall, the application of PCR assays for the detection of Salmonella from blood samples is an essential tool for the diagnosis and management of salmonellosis. Additional research is required to improve the sensitivity and specificity of these assays to detect Salmonella. Present findings also concluded that the most popular PCR techniques are multiplex PCR and RT-PCR for the detection of Salmonella may be due to their excellent sensitivity, specificity, and effectiveness.
The overall findings concluded that the most effective diagnostic approach for Salmonella infections should be chosen with a balanced assessment of efficiency, affordability, and accessibility. Real-time PCR emerges as a potent and effective technology for Salmonella detection, providing speedy and reliable findings by monitoring DNA amplification in real-time. While the initial cost and requirement for specialized equipment may be hurdles, the scalability of RT-PCR makes it well-suited for large-scale screening campaigns, contributing to its long-term cost-effectiveness. However, the context of the healthcare setting should not be overlooked. The simplicity and cost-effectiveness of traditional PCR for Salmonella identification remain important in resource-poor locations where infrastructure and understanding may be lacking. As technology advances, and with an emphasis on adapting diagnostic approaches to various situations, a holistic strategy that addresses both efficiency and accessibility becomes increasingly important.
ACKNOWLEDGMENTS
All the authors are thankful to GLA University, Mathura, UP, India for paying the article processing fee of this manuscript.
AUTHOR CONTRIBUTION
JD, SG and SD made a significant contribution to this review work. JD: data collection; writing-original draft; writing correction and editing. SG: supervision; project administration; investigation; validation and review. SD: concept making and methodology; formal analysis; investigation; editing and submission.
FINANCIAL SUPPORT
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
CONFLICTS OF INTEREST
The authors report no financial or any other conflicts of interest in this work.
ETHICAL APPROVALS
This study does not involve experiments on animals or human subjects.
DATA AVAILABILITY
All data generated and analyzed are included in this research article.
USE OF ARTIFICIAL INTELLIGENCE (AI)-ASSISTED TECHNOLOGY
The authors declares that they have not used artificial intelligence (AI)-tools for writing and editing of the manuscript, and no images were manipulated using AI.
PUBLISHER’S NOTE
All claims expressed in this article are solely those of the authors and do not necessarily represent those of the publisher, the editors and the reviewers. This journal remains neutral with regard to jurisdictional claims in published institutional affiliation.
REFERENCES
1. Carl AB. Encyclopedia of food microbiology. Cambridge, MA: Academic Press; 2014. Vol. 3, pp 349–52.
2. World Health Organization (WHO). Salmonella (non-typhoidal). Geneva, Switzerland: World Health Organization; [cited 2023 Nov 1]. Available from: https://www.who.int/news-room/fact-sheets/detail/salmonella-(non-typhoidal)
3. Ochiai RL, Acosta CJ, Danovaro-Holliday MC, Baiqing D, Bhattacharya SK, Agtini MD, et al. A study of typhoid fever in five Asian countries: disease burden and implications for controls. Bull World Health Organ. 2008;86(4):260–8. CrossRef
4. Pouzol S, Tanmoy AM, Ahmed D, Khanam F, Brooks WA, Bhuyan GS, et al. Clinical evaluation of a multiplex PCR for the detection of salmonella enterica serovars typhimurium and Paratyphi A from blood specimens in a high-endemic setting. Am J Trop Med Hyg. 2019;101(3):513–20. CrossRef
5. Centers for Disease Control and Prevention. Symptoms and treatment. Typhoid fever. Atlanta, Georgia: CDC; 2022.
6. NHS. Typhoid fever—vaccination—NHS choices. UK: NHS; 2021 [cited 2023 Sep 4]. Available from: http://www.nhs.uk/Conditions/Typhoid-fever/Pages/Prevention.aspx
7. Foddai ACG, Grant IR. Methods for detection of viable foodborne pathogens: current state-of-art and future prospects. Appl Microbiol Biotechnol. 2020;104(10):4281–8. CrossRef
8. Khokhar F, Pickard D, Dyson Z, Iqbal J, Pragasam A, John JJ, et al. Multiplex PCR assay to detect high risk lineages of Salmonella typhimurium and Paratyphi A. PLoS One. 2022;17(7):e0267805. CrossRef
9. Murphy MS. Management of bloody diarrhoea in children in primary care. BMJ. 2008;336(7651):1010–5. CrossRef
10. Larson DE, editor. Mayo clinic family health book: the ultimate home medical reference. New York, NY: William Morrow; 1993.
11. Smith M. Polymerase chain reaction (PCR). Bethesda, MD: Genome. gov.; 2023. Available from: https://www.genome.gov/genetics-glossary/Polymerase-Chain-Reaction
12. Ganesan V, Harish BN, Menezes GA, Parija SC. Detection of Salmonella in blood by PCR using iroB gene. J ClinDiagn Res. 2014;8(11):DC01–3. CrossRef
13. Kasturi KN, Drgon T. Real-time PCR method for detection of Salmonella spp. in environmental samples. Appl Environ Microbiol. 2017;83(14):e00644–17. CrossRef
14. Ding T, Suo Y, Zhang Z, Liu D, Ye X, Chen S et al. A multiplex RT-PCR assay for S. aureus, L. monocytogenes, and Salmonella spp. detection in raw milk with pre-enrichment. Front Microbiol. 2017;8:989. CrossRef
15. Levine MM, Grados O, Gilman RH, Woodward WE, Solis-Plaza R, Waldman W. Diagnostic value of the Widal test in areas endemic for typhoid fever. Am J Trop Med Hyg. 1978;27(4):795–800. CrossRef
16. Parry CM, Hien TT, Dougan G, White NJ, Farrar JJ. Typhoid fever. N Engl J Med. 2002;347(22):1770–82. CrossRef
17. Wain J, Pham VB, Ha V, Nguyen NM, To SD, Walsh AL et al. Quantitation of bacteria in bone marrow from patients with typhoid fever: relationship between counts and clinical features. J Clin Microbiol. 2001;39(4):1571–6. CrossRef
18. Wain J, Diep TS, Bay PV, Walsh AL, Vinh H, Duong NM et al. Specimens and culture media for the laboratory diagnosis of typhoid fever. J Infect Dev Ctries. 2008;2(6):469–74. CrossRef
19. Zhou L, Pollard AJ. A fast and highly sensitive blood culture PCR method for clinical detection of Salmonella enterica serovar typhimurium. Ann Clin Microbiol Antimicrob. 2010;9:14. CrossRef
20. Mullis KB. Facts. Nobel Prize.org. Nobel Prize Outreach AB 2023. [cited 2023 Dec 6]. Available from: https://www.nobelprize.org/prizes/chemistry/1993/mullis/facts/
21. Valones MAA, Guimarães RL, Brandão LA, de Souza PR, de Albuquerque Tavares Carvalho A, Crovela S. Principles and applications of polymerase chain reaction in medical diagnostic fields: a review. Braz J Microbiol. 2009;40(1):1–11. CrossRef,24031310.
22. Jensen B. Benefits of PCR testing for infectious diseases. Salt Lake City, UT: Bio fire diagnostics; 2021. Available from: https://www.biofiredx.com/blog/benefits-pcr-testing-infectious-diseases/
23. NIH, US. Web archive Retrieved from the Library of Congress. Bethesda, MD: National Human Genome Research Institute; [cited 2023 Dec 6]. Available from: https://www.genome.gov/genetics-glossary/Genetic-Marker
24. Garibyan L, Nidhi A. Polymerase chain reaction. J Investig Dermatol. 2013;133(3):1–4. CrossRef
25. Higginson EE, Nkeze J, Permala-Booth J, Kasumba IN, Lagos R, Hormazabal JC, et al. Detection of Salmonella typhimurium in bile by quantitative real-time PCR. Microbiol Spectr. 2022;10(3):e0024922. CrossRef
26. Ali A, Haque A, Haque A, Sarwar Y, Mohsin M, Bashir S et al. Multiplex PCR for differential diagnosis of emerging typhoidal pathogens directly from blood samples. Epidemiol Infect. 2009;137(1):102–7. CrossRef
27. Song JH, Cho H, Park MY, Na DS, Moon HB, Pai CH. Detection of Salmonella typhimurium in the blood of patients with typhoid fever by polymerase chain reaction. J Clin Microbiol. 1993;31(6):1439–43. CrossRef
28. Massi MN, Shirakawa T, Gotoh A, Bishnu A, Hatta M, Kawabata M. Rapid diagnosis of typhoid fever by PCR assay using one pair of primers from flagellin gene of Salmonella typhimurium. J Infect Chemother. 2003;9(3):233–7. CrossRef
29. Prakash P, Mishra OP, Singh AK, Gulati AK, Nath G. Evaluation of nested PCR in diagnosis of typhoid fever. J Clin Microbiol. 2005;43(1):431–2. CrossRef
30. Prabagaran SR, Kalaiselvi V, Chandramouleeswaran N, Deepthi KNG, Brahmadathan KN, Mani M. Molecular diagnosis of Salmonella typhimurium and its virulence in suspected typhoid blood samples through nested multiplex PCR. J Microbiol Methods. 2017;139:150–4. CrossRef
31. Zhou L, Jones C, Gibani MM, Dobinson H, Thomaides-Brears H, Shrestha S, et al. Development and evaluation of a blood culture PCR assay for rapid detection of Salmonella paratyphi A in clinical samples. PLoS One. 2016;11(3):e0150576. CrossRef
32. Park SH, Ricke SC. Development of multiplex PCR assay for simultaneous detection of Salmonella genus, Salmonella subspecies I, Salm. Enteritidis, Salm. Heidelberg and Salm. Typhimurium. J Appl Microbiol. 2015;118(1):152–60. CrossRef
33. Teh CSJ, Lau MY, Chong CW, Ngoi ST, Chua KH, Lee WS, et al. One-step differential detection of Salmonella entericaserovar typhimurium, serovar Paratyphi A and other Salmonella spp. by using a quadruplex real-time PCR assay. J Microbiol Methods. 2021;183:106184. CrossRef
34. Muttiullah F, Khan FM, Abbas FI, Shamim S. Characterization of different molecular markers for identification of Salmonella entericaserovar typhimurium in Pakistani population. J Bioresour Manag. 2017;4(4):4. CrossRef
35. Arkali A, Çetinkaya B. Molecular identification and antibiotic resistance profiling of Salmonella species isolated from chickens in eastern Turkey. BMC Vet Res. 2020;16(1):205. CrossRef
36. Malewski T, Malewska A. Rutkowski R. RT-PCR technique and its applications. State-of the-art. J Anim Feed Sci. 2003;12(3):403–16. CrossRef
37. Jordan JJ. Real-time detection of PCR products and microbiology. Trends Guide. 2000;8:61–6.
38. Tennant SM, Toema D, Qamar F, Iqbal N, Boyd MA, Marshall JM, et al. Detection of Typhoidal and Paratyphoidal Salmonella in blood by real-time polymerase chain reaction. Clin Infect Dis. 2015;61(4):S241–50. CrossRef.