INTRODUCTION
Cancer is a deadly disease caused by abnormal cell growth in certain parts of the body, having the ability to grow improperly, and spread to other parts of the body. According to the World Health Organization, around 4.4 million women died due to cancer in 2020, with 25% being caused by breast cancer (WHO, 2023). In contrast, cervical cancer affects women more frequently than any other form in 23 countries, with a total of 604,000 cases (IARC, 2022). Cancer treatment can be performed through therapy or medical appointments, such as surgery, radiotherapy, and chemotherapy (Hosseinzadeh et al., 2017). This method aims to suppress the growth and even destroy cancer cells, but this therapy will also damage healthy cells. In addition, the high cost of therapeutics and the success is not optimal. Therefore, herbal treatment approaches are in great demand in a way that the search for anticancer secondary metabolites derived from plants is fundamental.
The genus Araucaria belongs to the family Araucaria. It consists of 19 species, and 8 species of them contain biflavonoids, including Araucaria cookii (Ilyas et al., 1977), Araucaria excelsa (Aslam et al., 2013) Araucaria hunsteinii (Agusta et al., 2022), Araucaria rulei F. Muell, Araucaria angustifolia, Araucaria columnaris, Araucaria araucana, Araucaria bidwilli Hook, and Araucaria cunninghamii Mudie (Frezza et al., 2020). Biflavonoids from several Araucaria species, such as amentoflavone, agathisflavones, cupresuflavones, robusta flavones, and hinokiflavones, play a range of biological activities (Frezza et al., 2020). Considering the variety of biflavonoid content and its activities, it is essential to investigate the biflavonoid content of plants belonging to the genus Araucaria. Subsequently, Araucaria plants are found in the southern hemisphere, such as New Guinea, Argentina, Australia, Chile, Brazil, Norfolk Island, and New Caledonia (Frezza et al., 2020). In addition, plants of this genus also are found in Indonesia, such as A. bidwilli, A. cunninghamii, Araucaria heterophylla, and A. columnaris (Frost. f.) Hookf, and A. hunsteinii K. Schum. grows in the Bogor and Cibodas Botanical Gardens.
Biflavonoid is one of the secondary metabolites with high development potential as an anticancer agent (Xie et al., 2021). They are dimers of flavonoids linked by oxidation through C-O-C or C-C bonds (Menezes and Campos, 2021). One of the known biflavonoid compounds, 7.5?-di-O-methylrobustaflavone inhibition concentration 50 (IC50 11.89 μM) has been reported to inhibit the Michigan Cancer Foundation-7 (MCF-7) cell line (breast cancer cells) possibly through the mitochondrial regulatory pathway (Xie et al., 2021), while 4′, 4?′- di-O-methyl amentoflavone or isoginkgetin (IC50 8.38 ± 0.63 μM) can induce apoptosis of HeLa cancer cells (cervical cancer cells) (Li et al., 2019). Biflavonoids not only have an anticancer effect but also have antibacterial (Deforest et al., 2014), antidiabetic and anti-inflammatory (Ayepola et al., 2014), antiviral (Menezes and Campos, 2021), antioxidant, and antifungal (Yu et al., 2017). A previous study on the leaves of A. hunsteinii K. Schum. succeeded in isolating two bilfavonoids (Agusta et al., 2022) namely 4′,7,7?-triO-methylcupressuflavone and 4?′,7,7?- tri-O-methylagathisflavone. Due to the lack of information on secondary metabolites, specifically biflavonoids from the Indonesian plant A. hunsteinii K. Schum. plant, this study aimed to isolate the additional biflavonoids present in the leaves of A. hunsteinii and evaluate their inhibitory activity against the MCF-7 and HeLa cancer cells, and by examining the structure-activity relationship (SAR) of all isolated biflavonoids.
MATERIAL AND METHODS
Material, reagents, and instrumentation
The materials used in this research are acetone fractions of A. hunsteinii leaves free from chlorophyll and tannins that is resulted by Agusta et al. (2022), the technical grade organic solvent, such as acetone, methanol (MeOH), ethanol (EtOH), dichloromethane (CH2Cl2), and chloroform (CHCl3). The separation process used a chromatography method with various support solids such as Sephadex LH-20 (GE Healthcare), silica gel 60 Merck 60 (0.040–0.063 mm) and (0.063–0.2 mm), and GF254 Merck thin layer chromatography (TLC-preparative). The isolated compounds were characterized according to the method described by Agusta et al. (2022). In addition, MCF-7 (ATTC HTB 22) and HeLa cancer cells (ATTC CCL 2) cancer cells were subjected to the cytotoxic anticancer test using the 3-(4,5- dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT) reagent. Both cells were grown in 24 well microplates at a concentration of 5,000 cells in 100 μl of growth medium (D-MEM, Roswell Park Memorial Institute 1640, 5% fetal bovine serum, and Penicillin 100 U/ml, Streptomycin 100 ug/ml). These two cells were from the IPB University’s Primate Research Center. HeLa (ATTC CCL-2) cell is obtained from adenocarcinoma-affected human uterine and cervix epithelial cells, whereas MCF-7 (ATTC HTB 22) cell is derived from human breast and mammary gland epithelial cells. They were kept in liquid nitrogen’s vapor phase under frozen storage conditions.
Fractionation and isolation of A. hunsteinii leaves fractions
The 5.0 and 10.0-g acetone fractions were separated using Sephadex LH-20, yielding 31 fractions (F1–F31) and 17 fractions (G1–G17), respectively. Biflavonoids are expected to be present in the F1–F14 and G1–G11 fractions of the TLC analysis using the eluent of CHCl3: MeOH (19:1). The yellow dots that show up on TLC after exposure to Ce(SO4)2 solution are biflavonoids. All fractions were purified using silica gel column chromatography (CC) and TLC-preparative. The purification in CC silica gel was performed on the F3, F8, F10, G5, G6, G7, and G8. The F8 fraction was eluted with CH2Cl2:MeOH (65:1) eluent and further purified with CH2Cl2:EtOH (50:1) eluent to obtain 1 (16.8 mg). The F3 fraction (91.9 mg) was eluted with CH2Cl2 eluent with increasing polarity and obtained 5 (26.5 mg). The combined fractions F10 (75.0 mg), G5 and G6 (243 mg), and G7–G8 (174.5 mg) were eluted with CH2Cl2:EtOH (30:1) eluent and afforded 2 (35.3 mg), 3 (20.4 mg) and 4 (21.8 mg). The elusion results were observed under ultraviolet (UV) light at 254 and 366 nm, and the fraction was sprayed with a solution of Ce(SO4)2. Schematic diagrams of the biflavonoids isolation can be seen in Figures S1 and S2. All compounds were characterized by using spectrophotometers of 1D and 2D nuclear magnetic resonance (NMR), UV-visible, infrared (IR), and liquid chromatography-mass spectrometry tandem mass spectrometry (LC-MS/MS), and identified as 7,4?′-di-O-methylcupressuflavone (1), 7-O- methylcupressuflavone (2), 4′, 4?′-di-O-methyl amentoflavone or isoginkgetin (3), 7,7?-di-O- methylagathisflavone (4) and 7,4′,7?,4?′-tetra-O-methylcupressuflavone (5). The structure of the five compounds is shown in Figure 1.
Cytotoxicity assay
The MTT method against MCF-7 cell lines is described in the protocol by Sasikala et al. (2020) to perform the cytotoxicity test. After 50% confluency, the cells were grown with 5,000 cells each containing well in 100 ml of the growth medium, the different concentrations of isolated compounds (Table S1) and 10 μl of MTT reagent (5 mg/ml) was added, and then 4 hours at incubated at room temperature (37°C) until the formazan crystal was formed.
The MTT method against HeLa cell lines is described in the protocol by Nawaz et al. (2021). The cells were grown with a concentration of 5,000 cells, each containing well in 100 ml of growth medium after 50% confluency. The different concentrations of isolated compounds (Table S2) and 10 μl of MTT reagent (5 mg/ml) were added, and then incubated for 4 hours at room temperature (37°C). The formed formazan crystals were dissolved in EtOH and then monitored with a spectrophotometric plate reader at 595 nm. The assay had been conducted in three replicates. The percentage of cell inhibition was calculated using the formula:
The IC50 value of the sample is obtained from the linear regression equation y = a× + b, where x is the concentration of the sample and y is the % inhibition of the sample. The statistical evaluation was given in all IC50 values. The morphology of the conserved MCF-7 and HeLa cells was observed under an inverted microscope with 400 times magnification.
RESULTS AND DISCUSSION
This study was performed by isolating five biflavonoids. There are 7,4?′-di-O- methylcupressuflavone (1), 7-O-methylcuppresuflavone (2), 4′,4?′-di-O-methylamentoflavone or isoginkgetin (3), 7,7?-di-O-methylagathisflavone (4) and 7,4′,7?,4?′-tetra-O-methylcupressuflavone (5). The structure of the five compounds is shown in Figure 1. These compounds were firstly discovered in the leaves of A. hunsteinii. Compound 1 is a new compound that has never been reported before and the first to be isolated from the Araucaria genus. Moreover, 2 to 5 have been found in other species of the family Araucaria. Compound 5 has been found in A. cunninghamii, A. rulei, A. columnaris, Wollemia nobilis, and Agathis ovata. Meanwhile, 2 have been found in A. bidwilli, Agathis atropurpurea, A. ovata, Araucaria australis, Araucaria robusta, and Agathis alba. 3 were found only in the genus Araucaria, there are A. angustifolia and A. bidwilli, while 4 have been reported in A. alba, A. robusta, A. ovata, A. bidwilli, and A. columnaris (Frezza et al., 2020; Khan et al., 1972).
Identification of biflavonoids
The basic structure of biflavonoids has 30 carbon atoms. The 1H NMR and 13C-NMR data reveal various properties of these compounds. On the other hand, biflavonoids showed two distinct signals in their 1H-NMR spectra at δH 13.00 ppm, indicating the presence of chelation between protons and carbons of hydroxyl groups and carbonyl groups. 13C-NMR spectra of biflavonoids showed 2 peaks at δC 182.00 ppm, and biflavonoids also have 10 characteristic peaks at δC 160.0–166.0 ppm. The protons of the methoxy group of biflavonoids are known from the proton chemical shift value of δH 3.75–3.85 ppm with a higher 13C-NMR shift value than the carbon in the hydroxyl group because the carbon attached to the hydroxyl group is more shielded than the methoxy group. In the 1H-NMR spectrum, 1 has 9 signals, 3 has 14 signals, and 4 has 12 signals representing 22 protons. In addition, 2 has 11 signals of 1H-NMR representing 20 protons, while 5 has 7 signals representing 26 protons. Moreover, the 13C-NMR spectrum of 1, 3, and 4 has 28 signals representing 32 carbons, while 2 has 26 signals representing 31 carbons and 5 has 15 signals representing 34 carbons.
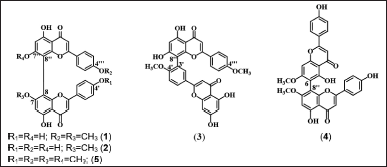 | Figure 1. Structure of compounds 1–5 isolated from the leaves of A. hunsteinii. [Click here to view] |
Cupressuflavone derivatives (1, 2, and 5)
7,4?′-Di-O-methylcupressuflavone (1)
Compound 1 was acquired as pale-yellow amorphous powder with the Rf value of 0.39 (CHCl3:MeOH = 19:1). The UV data (CH3OH) showed the absorbance at wavelength (λmax) of 273 nm that showed benzoyl chromophore and 329 nm that showed cinnamoyl chromophore from flavonoids structure. The IR data (KBr) showed that 1 have several functional groups such as hydroxyl groups at wave number (νmax) of 3,304 cm−1, carboxyl group (C=O) at 1,661 cm−1, C=C aromatic at 1,572 cm−1, also ether groups (C–O) at 1,278–1,205 and 1,098–1,066 cm−1.
The 1H NMR (CD3OD, 500 MHz) data of 1 showed three doublet signals of two aromatics rings which para substituted δ 7.46 (1H, d, J = 8.9 Hz, H-2′, -2?′, -6′, and -6?′), 6.87 (1H, d, J = 9.0 Hz, H-3?′ and -5?′), 6.73 (1H, d, J = 8.7 Hz, H-3′ and -5′); four singlet aromatic signals δ 6.69 (1H, s, H-6), 6.61 (1H, s, H-3 and H-3?), 6.46 (1H, s, H-6?); and two signals represent two methoxy groups at δ 3.84 (3H, s, 7-OCH3), 3.78 (3H, s, 4?′-OCH3). The 13C NMR (CD3OD, 126 MHz) data displayed: 2 signals of hydroxy groups chelated carbonyls δ 184.4 (C, C-4), 184.2 (C, C-4?); 10 signals of oxyaril carbons δ 166.4 (C, C-2), 165.5 (C, C-2?), 165.3 (C, C-7), 164.3 (C, C-4?′), 164.1 (C, C-7’’), 163.8 (C, C-5), 162.9 (C, C-5?), 162.8 (C, C-4’), 156.8 (C, C-9), 156.1 (C, C-9?); 13 signals represent 18 sp2 aromatic carbons δ 129.2 (CH, C-2′ and -6′), 128.8 (CH, C-2?′ and -6?′), 124.2 (C, C-1?′), 123.0 (C, C-1′), 116.9 (CH, C-3′ and -5′), 115.5 (CH, C-3?′ and -5?′), 105.9 (C, C-10), 105.5 (C, C-10’’), 103.9 (CH, C-3?), 103.4 (CH, C-3), 101.0 (C, C-8), 99.9 (C, C-8?), 99.8 (CH, C-6?), 96.4 (CH, C-6); and two signals of methoxy carbons δ 56.9 (CH3, 7-OCH3), 56.0 (CH3, 4?′-OCH3). LC-MS/MS dimethyl sulfoxide (DMSO): LC rt 10.50 minutes, electrospray ionization mass spectrometry (ESI/MS) m/z calculated for C32H22O10: 567.1291 [M+H]+, 552.1012, 535.0977, 449.0840, 435.0678, 390.0707, 346.0450, 297.0744, 267.0640, 258.0148, 228.0047, 135.0441, 121.0285. The UV, IR, and NMR spectra also the LC-MS chromatogram of compound 1 can be seen in Figures S3–S10.
7-O-methylcupressuflavone (2)
Compound 2 was achieved as yellow amorphous powder with the Rf value of 0.37 (CHCl3:MeOH = 19:1), The UV data (CH3OH) showed the absorbance at λmax 273 nm (benzoyl chromophore) and 330 nm (cinnamoyl chromophore) which indicates 2 has a flavonoid structure. The IR data (KBr) showed that 2 have several functional groups such as hydroxyl groups at νmax 3,329 cm−1, C=C aromatic at 1,446 cm−1, and ether group (C–O) at 1,240 cm−1.
The 1H NMR (Acetone-d6, 500 MHz) data of 2 showed: two signals of hydroxy groups chelated carbonyl δ 13.38 (1H, s, 5-OH), 13.22 (1H, s, 5?-OH); four doublet signals of two aromatics rings which para substituted δ 7.60 (1H, d, J = 8.8 Hz, H-2′ and -6′), 7.53 (1H, d, J = 8.8 Hz, H-2?′ and -6?′), 6.86 (1H, d, J = 8.8 Hz, H-3′ and -5′), 6.85 (1H, d, J = 8.8 Hz, H-3?′ and -5?′); three singlet aromatic signals δ 6.68 (1H, s, H-3 and -6), 6.67 (1H, s, H-3?), 6.53 (1H, s, H-6?); and a signal represent a methoxy group at δ 3.88 (3H, s, 7-OCH3). The 13C NMR (Acetone-d6, 126 MHz) data displayed: two signals of hydroxyl groups chelated carbonyls δ 183.6 (C, C-4), 183.4 (C, C-4?); 10 signals of oxyaril carbons δ 165.3 (C, C-2), 164.8 (C, C-2?), 164.8 (C, C-7), 163.8 (C, C-5), 163.1 (C, C-5?), 162.9 (C, C-7?), 161.9 (C, C-4′), 161.8 (C, C-4?′), 156.2 (C, C-9?), 155.8 (C, C-9); 14 signals of sp2 aromatic carbons δ 129.0 (CH, C-2′ and -6′), 128.8 (CH, C-2?′ and -6?′), 123.0 (C, C-1′), 123.0 (C, C-1?′), 116.9 (CH, C-3′ and -5′), 116.7 (CH, C-3?′ and -5?′), 105.7 (C, C-10), 105.5 (C, C-10?), 103.7 (CH, C-3), 103.6 (CH, C-3?), 100.1 (C, C-8), 99,6 (CH, C-6?), 99.5 (C, C-8?), 96.1 (CH, C-6); and a signal of methoxy carbons δ 56.8 (CH3, 7-OCH3). LC-MS/MS (Acetone): LC rt 9.34 minutes, ESI/MS m/z calculated for C31H20O10: 553.3317 [M+H]+, 462.8165, 391.3195, 371.3495, 352.2081, 279.1895, 195.1081, 141.9752. The UV, IR, and NMR spectra also the LC-MS chromatogram of compound 2 can be seen in Figures S11–S18.
4′,4?′,7,7?-Tetra-O-methylcupressuflavone (5)
Compound 5 was gained as yellow amorphous powder with the Rf value of 0.85 (CHCl3:MeOH = 19:1), The UV data (CH3OH) showed the absorbance at λmax 273 nm (benzoyl chromophore) and 326 nm (cinnamoyl chromophore) which indicates 5 has a flavonoid structure. The IR data (KBr) showed that 5 have several functional groups such as carbonyl groups at νmax 1,612 cm−1, C=C aromatic at 1,437 cm−1, also ether groups at 1,259–1,181 cm−1 and 1,099–1,039 cm−1.
The 1H NMR (CDCl3, 500 MHz) data of 5 showed: a signal of hydroxy group chelated carbonyl δ 13.22 (1H, 5 and 5?-OH); two doublet signals of two aromatics rings which para substituted δ 7.43 (1H, d, J = 8.9 Hz, H-2′, -2?′, -6′, and -6?′), 6.86 (1H, d, J, = 8.9 Hz, C-3′, -3?′, -5’ and -5?′); two singlet aromatic signals δ 6.60 (1H, s, C-3 and -3?), 6.59 (1H, s, C-6 and -6?); and two signals represent a methoxy group at δ 3.82 (3H, s, 7- and 7?-OCH3), 3.80 (3H, s, 4′- and 4?′-OCH3). The 13C NMR (CDCl3, 126 MHz) data displayed: a signal of hydroxy group chelated carbonyls δ 183.1 (C, C-4 and -4?); 5 signals of oxyaril carbons δ 164.1 (C, C-2 and -2?), 163.5 (C, C-4′ and -4’’’), 162.8 (C, C-7 and -7?), 162.7 (C, C-5 and C-5?), 154.8 (C, C-9 and -9?); 7 signals of sp2 aromatic carbons δ 127.8 (CH, C-2′, -2?′, -6′, and -6?′), 123.4 (C, C-1′ and -1?′), 114.7 (CH, C-3′, -3?′, -5’, and -5?′), 105.4 (C, C-10 and -10?), 103.7 (CH, C-3 and -3?), 99.6 (C, C-8 and -8?), 95.44 (CH, C-6 and -6?); and two signals of methoxy carbons δ 56.4 (CH3, 7 and 7?-OCH3), 55.7 (CH3, 4′ and 4?′-OCH3). LC-MS/MS (CD2Cl2): LC rt 13,02 minutes, ESI/MS m/z calculated for C34H26O10: 595.1628 [M+H]+, 580.1370, 549.1192, 463.1029, 430.0692, 403.0809, 374.0784, 359. 0567, 297.0768, 284.0692, 135.0455. The UV, IR, and NMR spectra also the LC-MS chromatogram of compound 5 can be seen in Figures S35–S42.
Cupressuflavone is a flavone dimer binding at C8 and C8? as shown by evidence from heteronuclear singular quantum coherence (HSQC) and heteronuclear multiple bond coherence (HMBC). The HSQC spectrum gives information about the correlation of a bond between the corresponding proton and carbon. At the same time, the HMBC showed a correlation between proton and carbon separated by two to three bonds. Both C8 and C8? correlate with H6 and H6? respectively, but do not bind with protons (quaternary carbons). HMBC correlation also can determine the location of methoxy groups. The proton of methoxy groups in 1 showed HMBC correlation with C4?′ and C7, while 5 showed correlations with C4′, C4?′, C7, and C7?. Compound 2 showed an HMBC correlation between the proton of the methoxy functional group and C7. HSQC and HMBC correlation of the cupressuflavone group can be seen in Figure 2.
LC-MS/MS analysis confirmed that the NMR data of 1, 2, and 5 are biflavonoids and not flavonoid monomers since the molecular weight of 1, 2, and 5 is twice that of flavonoids. The molecular weights of 1, 2, and 5 are m/z 567.1291 [M+H]+ (C32H22O10); 553.3317 [M+H]+ (C31H20O10); and 595.1628 [M+H]+ (C34H26O10) respectively. In 5 analyses, the NMR data showed that half of the biflavonoids were nearly symmetrical. However, the results of the LC-MS/MS measurement of 5 showed that the compound has a molecular weight twice higher than the flavonoids. Therefore 5 is a large compound consisting of two flavonoids with a symmetrical structure. The fragmentation pathways of the cupressuflavone groups are shown in Figure 3. The flavone C-ring was cleaved at fragment m/z 567 becomes 449 at 1, 553 becomes 391 and 463 becomes 359 at 2, also 549 becomes 403 and 430 becomes 374 at 5. The termination of the functional group occurs in the fragment m/z 553 becomes 463 and 297 becomes 255 at 2, and 521 becomes 463 at 5. Similarly, the cleavage of the hydroxyl group also occurs in fragments m/z 553 becomes 463 at 2 and 521 becomes 463, and 374 becomes 359 at 5 (Nakata et al., 2018; Zhang et al., 2011).
Amentoflavone derivative (3)
4′,4?′-Di-O-methylamentoflavone (Isoginkgetin, 3)
Compound 3 was afforded as pale-yellow amorphous powder with the Rf value of 0.27 (CHCl3:MeOH = 19:1), The UV data (CH3OH) showed the absorbance at λmax 270 nm (benzoyl chromophore) and 331 nm (cinnamoyl chromophore) which indicates 3 has a flavonoid structure The IR data (KBr) showed 3 have several functional groups such as carbonyl groups at νmax 1,653 cm−1, C=C aromatic at 1,599 cm−1, and ether group at 1,279–1,261 cm−1.
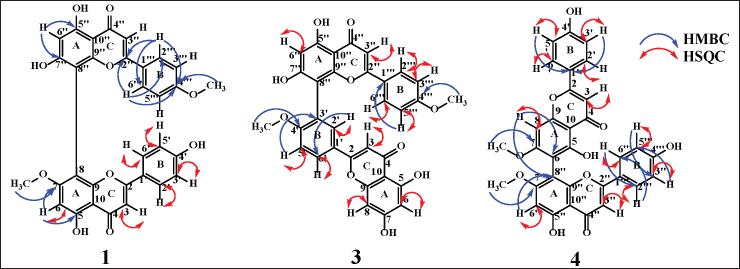 | Figure 2. HSQC and HMBC key correlation of 1 (cupressuflavone), 3 (amentoflavone), and 4 (agathisflavone). [Click here to view] |
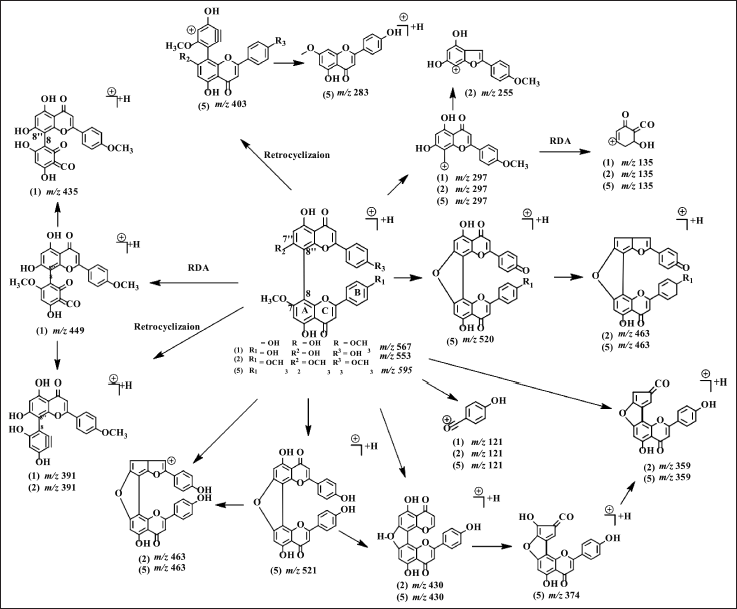 | Figure 3. Proposed MS fragmentation scheme of cupressuflavones (1, 2, and 5). [Click here to view] |
The 1H NMR (DMSO-d6, 500 MHz) data of 3 showed two signals of hydroxy groups chelated carbonyl δ 13.18 (1H, 5?-OH), 12.84 (1H, 5-OH); six doublet signals of two aromatics rings which para substituted δ , 8.11 (1H, dd, J = 2.4, 8.7 Hz, H-6′), 8.01 (1H, d, J = 2.4 Hz, H-2′), 7.44 (1H, d, J = 8.8 Hz, H-3?′ and -5?′), 7.29 (1H, d, J = 9.0 Hz, H-5′), 6.41 (1H, d, J = 2.0 Hz, H-8), 6.12 (1H, d, J = 2.0 Hz, H-6); four singlet aromatic signals δ 6.85 (1H, s, H-3), 6.77, (1H, s, H-3?) 6.64 (1H, d, J = 8.8 Hz, H-2?′ and -6?′), 6.59 (1H, s, C-6?); and two signals represent two methoxy groups at δ 3.76 (3H, s, 4?′-OCH3), 3.71 (3H, s, 4′-OCH3). The 13C NMR (DMSO-d6, 126 MHz) data displayed: two signals of hydroxy groups chelated carbonyls δ 182.4 (C, C-4?), 181.8 (C, C-4); nine signals of oxyaril carbons δ 164.4 (C, C-7), 164.0 (C, C-2?), 163.3 (C, C-2), 162.6 (C, C-5), 161.5 (C, C-5? and -7?), 161.3 (C, C-4?′), 160.4 (C, C-4′), 157.5 (C, C-9), 153.5 (C, C-9?); 16 signals of sp2 aromatic carbons δ 130.9 (CH, C-2′), 128.4 (CH, C-2?′ and -6?′), 128.2 (CH, C-6′), 122.6 (C, C-1’), 121.3 (C, C-1?′), 121.2 (C, C-3’), 115.9 (CH, C-3?′ and -5?′), 111.9 (CH, C-5′), 104.7 (C, C-10?), 104.1 NMR spectra also the LC-MS chromatogram of compound 3 can be seen in Figures S19–S26.
Amentoflavone has a dimer bond at quaternary carbons C3′ and C8?, confirmed by HSQC and HMBC data (Fig. 2). The 3′ carbon showed an HMBC correlation with H5′ while carbon 8? exhibited HMBC correlation with H6?. The proton of methoxy groups in 3 has HMBC correlation with C4′, and 4?′ which indicates methoxy groups at C4′ and 4?′. HSQC and HMBC correlation of 3 can be seen in Figure 2. LC-MS/MS fragmentation analysis confirms the NMR data of 3 is biflavonoid. The molecular weight of 3 is m/z 567.1318 [M+H]+ (C32H22O10). Cleavage of dimer bond in amentoflavone-type biflavonoids occurs at fragment m/z 567 to 297. The flavone C-ring was cleaved at fragment m/z 521 becomes 403 (Nakata et al., 2018; Zhang et al., 2011), the MS fragmentation of 3 can be seen in Figure 4.
Agathisflavone derivative (4)
7,7?-Di-O-methylagathisflavone (4)
Compound 4 was obtained as yellow amorphous powder with the Rf value of 0.35 (CHCl3:MeOH = 19:1), The UV data (CH3OH) showed the absorbance at λmax 273 nm (benzoyl chromophore) and 330 nm (cinnamoyl chromophore) which indicates 4 has a flavonoid structure. The IR data (KBr) showed 4 have several functional groups such as hydroxyl groups at νmax 3,329 cm−1, C=C aromatic at 1,446 cm−1, and ether group at 1,240 cm−1.
The 1H NMR (Acetone-d6, 500 MHz) data of 4 showed: two signals of hydroxy groups chelated carbonyl δ 13.29 (1H, s, 5-OH), 13.26 (1H, s, 5?-OH); five doublet signals of two aromatics rings which para substituted δ 8.03 (1H, d, J = 8.8 Hz, H-2′ and -6′), 7.61 (1H, d, J = 8.8 Hz, H-2?′ and H-6?′), 7.07 (1H, d, J = 8.3 Hz, H-3′ and -5′), 6.99 (1H, d, J = 2.0 Hz, H-8) 6.85 (1H, d, J = 8.9 Hz, H-3?′ and -5?′); three singlet aromatic signals δ 6.77 (1H, s, H-3), 6.66 (1H, s, H-3?), 6.57 (1H, s, H-6?); and a signal represent a methoxy group at δ 3.88 (3H, 7- and 7?-OCH3). The 13C NMR (Acetone-d6, 126 MHz) data displayed: two signals of hydroxy groups chelated carbonyls δ 182.7 (C, C-4?), 182.4 (C, C-4); 10 signals of oxyaril carbons δ 164.5 (C, C-2), 164.3 (C, C-2?), 163.8 (C, C-7?), 163.7 (C, C-7), 162.4 (C, C-5?), 161.4 (C, C-4′), 161.2 (C, C-4?′), 159.5 (C, C-9), 157.9 (C, C-5), 154.4 (C, C-9?); 13 signals of sp2 aromatic carbons δ 128.5 (CH, C-2′ and -6′), 128.1 (CH, C-2’’’ and -6?′), 122.2 (C, C-1′ and -1?′), 116.1 (CH, C-3′ and -5′), 115.9 (CH, C-3?′ and -5?′), 105.1 (C, C-10), 104.6 (C, C-10?), 104.2 (C, C-6), 103.5 (CH, C-3), 102.7 (CH, C-3?), 100.3 (C, C-8?), 95.2 (CH, C-6?), 90.3 (CH, C-8); and a signal of methoxy carbons δ 55.9 (CH3, 7 and 7?-OCH3). LC rt 10.34 minutes, ESI/MS m/z calculated for C32H22O10: 567.1371[M+H]+, 521.0929, 447.1125, 431.0812, 405.1014, 389.0701, 361.0741, 332.0716, 285.0437, 242.0253, 186.0343, 121.0308. The UV, IR, and NMR spectra also the LC-MS chromatogram of compound 4 can be seen in Figures S27–S34.
Agathisflavone showed a dimer bond at quaternary carbons C6 and C8? from HSQC and HMBC data (Fig. 2). Based on the HMBC data, C6 shows HMBC correlation with H8, while C8? has an HMBC correlation with H6?. The proton of methoxy groups in 4 has HMBC correlation with C7 and 7?, indicating methoxy groups at C7 and 7?. The HSQC and HMBC correlation of 4 can be seen in Figure 2. LC-MS/MS analysis confirmed the NMR data of 4 is biflavonoid, and the flavone C-ring cleaved at fragment m/z 567 becomes 447. The cleavage of dimer bond in amentoflavone-type biflavonoids occurs at fragment m/z 567 becomes 297 and 361 becomes 285. The termination of the functional group occurs in the fragment m/z 447 becomes 405 and 431 becomes 361, and the cleavage of the hydroxyl group also occurs in fragments m/z 447 becomes 431 (Nakata et al., 2018; Zhang et al., 2011). The MS fragmentation scheme of 4 can be seen in Figure 5.
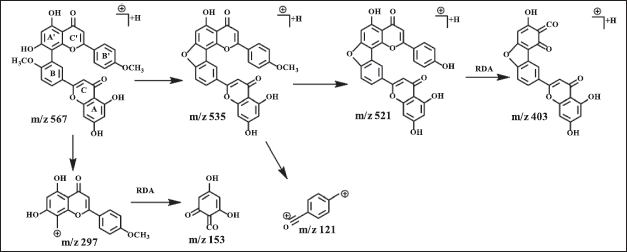 | Figure 4. Proposed MS fragmentation scheme of amentoflavone (3). [Click here to view] |
Cytotoxicity activity of biflavonoids against MCF-7 and HeLa cancer cells
The cytotoxicity assay against MCF-7 cancer cells
The cytotoxicity test for MCF-7 and HeLa cancer cells was performed at 1-5. The data can be Table S1 and S2. The cytotoxicity test of biflavonoids against MCF-7 and HeLa cancer cells respectively was evaluated qualitatively by observing cell morphology with an inverted microscope using a spectrophotometric plate reader at a wavelength of 595 nm. Figure 6 shows the morphology of MCF-7 cells with 400× magnification in the negative control, the positive control (Epirubicin-HCl) with a concentration of 31.25 ppm, and 2 with a concentration of 31.25 ppm. The cell morphology in the negative control (Fig. 6N1) showed cells growing on the surface of the plate (confluency close to 100%). The degree of confluency is defined as the percentage of the surface area covered by the cells (Haenel and Garbow, 2014). Living cells have well-defined membranes with clear, light- colored, and epithelial-shaped media. MCF-7 cells given a positive control (Fig. 6N2) and 2 at the same concentration (Fig. 6N3) showed confluent cell death, cell membrane boundaries were not visible, and the cell shape was irregular. However, based on this qualitative analysis, it needs to be ascertained whether the sample added to the cell enters entirely and thoroughly into the cell. According to Yeap et al. (2021), MCF-7 cells have polygonal cell shapes with clear and distinctive cell membrane boundaries, while dead cells have distorted figures. This is different from the characteristics of living cells shown in the negative control (Fig. 6N1), so the addition of compound 2 indicates a cell response.
This test was carried out using the MTT method, which is relatively faster, more sensitive, and accurate, and can be applied to various samples (van Tonder et al., 2015). Table 1 shows the IC50 values of cytotoxicity of the five compounds against the MCF-7 cancer cells. Moreover, the cytotoxicity test showed that 1, 2, and 3 are classified as very active, 4 as moderately active, and 5 as weakly active in inhibiting the growth of MCF-7 cells. According to the National Cancer Institute (NCI) of the United States, the cytotoxicity of the sample tested can be categorized as very active if the IC50 < 20 μM, moderate activity if IC50 ranged between 21 and 200 μM, weak activity if IC50 ranged between 201 and 500 μM and not-active if IC50 >500 μM (Damasuri et al., 2020).
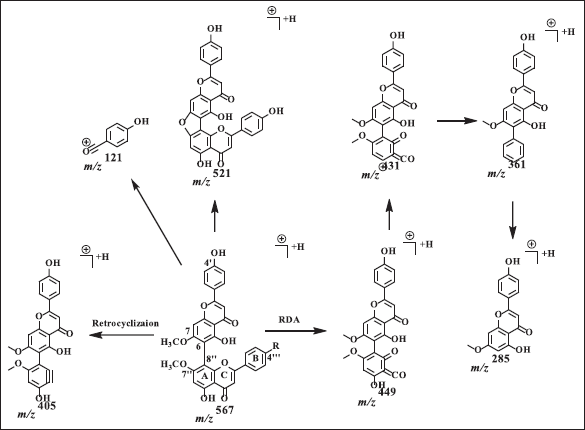 | Figure 5. Proposed MS fragmentation scheme of agathisflavone (4). [Click here to view] |
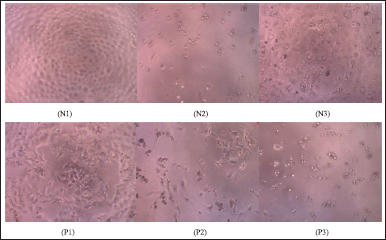 | Figure 6. The morphology of MCF-7 cells with 400× magnification on the negative control (N1), positive control at a concentration of 31.25 ppm (N2), and the addition of 2 at a concentration of 31.25 ppm (N3). The HeLa c cancer cells ell morphology of negative control (P1), positive control at a concentration of 31.25 ppm (P2), and the addition of 3 at a concentration of 31.25 ppm (P3). [Click here to view] |
 | Table 1. IC50 value of isolated compounds 1–5 against MCF-7 and HeLa cancer cells. [Click here to view] |
The cytotoxicity assay against HeLa cancer cells
Qualitative analysis of HeLa cancer cells was carried out by observing the morphology of cancer cells with an inverted microscope. The morphology of HeLa cancer cells in the negative control, positive control (Napradox-50), and compound 3 at a concentration of 31.25 ppm, respectively are shown in Figure 6. The cell morphology in the negative control (Fig. 6P1) shows that HeLa cancer cells containing viable organisms appear confluent, oval in shape, flat, light in color and have clear cell membrane boundaries. Cells supplemented with Napradox-50 at a concentration of 31.25 ppm (Fig. 6P2) showed that the cells were dead, indicated by irregular shape and black cells, but viable HeLa cancer cells were still visible. HeLa cancer cells with the addition of 3 at the same concentration (Fig. 6P3) only showed dead cells that were small and scattered. This suggests that adding 2 to MCF-7 cells and 3 to HeLa cancer cells indicates a cell response, which causes death in these cells.
The quantitative analysis using MTT assay against HeLa cancer cells showed that 2 and 3 are classified as very active, 1 and 4 compounds are moderately active, and 5 are inactive. The activity was categorized according to the NCI of the United States (Damasuri et al., 2020).
Comparison of biflavonoid activity against MCF-7 and HeLa cancer cells
As shown in Table 1, 2 and 3 are the most active compound to inhibit MCF-7 and HeLa cancer cells, respectively. The IC50 value of 2 was observed in the order of HeLa cancer cells
Many pathways have been proposed based on the literature review for the mechanism that prevents the action of three in cancer cell invasion. According to Yoon et al. (2006), 3 can control the synthesis of the metalloproteinase-9 matrix in B16F10 melanoma, MDA-MB-231 breast carcinomas, and HT1080 human fibrosarcoma cells by inhibiting NF-B (factor kappa B) and Akt (protein kinase B). In another study, Tsalikis et al. (2019) found that 3 directly inhibited caspase, trypsin, and chymotrypsin-like activities connected to the beta-1, 2, and 5 subunits of the 20S proteasome. Additionally, O’Brien et al. (2008) reported that 3 also inhibits pre-mRNA splicing in HeLa cancer cells. The pre-mRNA splicing is essential for the emergence and development of different types of cancer.
The amount and position of methoxy groups regulate the biological activity of compounds, as demonstrated by the methylated derivatives of the cupressuflavone 7,4′,7?-tri-O- methylcupressuflavone (A1) (Agusta et al., 2022), 1, 2, and 3 on MCF-7 and HeLa cancer cells (Table 1). A compound with mono-, di- and/or trimethyl groups (A1, 1, and 2) has a more excellent activity (respectively, IC50 of 91.74 ± 5.6, 11.54 ± 3.4, and 3.40 ± 0.3 μM) on MCF-7 than 5 (IC50 = 397.89 ± 28.6 μM). The same case for the cellular compounds HeLa cancer cells A1 and 2 shows a higher activity high with IC50 35.59 ± 1.3 and 1.42 ± 1.1 μM, respectively, than 5 (IC50 = 528.78 ± 40.0 μM). In addition, 5 shows a weaker activity for both cells. The existence of methoxy groups at C4′, C7?, and/or C4?′ positions appears to slightly reduce the activity. Methylations at position C7 on the cupressuflavone skeleton appear to be the most beneficial for inhibiting the MCF-7 and HeLa cancer cells’ activity because 2 were more active than A1, 1, and 5. According to a previous study, the minor difference in activity caused by the position and number of methoxy groups may be due to the ability to form hydrogen bonds to MCF-7 or HeLa cancer cells and changing polarity (Molfetta et al., 2004). Agathisflavone derivatives (4) are slightly more active inhibitors on both cells MCF-7 (115.39 ± 36.5 μM) and HeLa cancer cells (IC50 = 107.63 ± 37.3 μM) than 7,7?,4?′- tri-O- methylagathisflavone (A2) for MCF-7 (IC50 = 314.44 ± 25.0 μM) and HeLa cancer cells (IC50 = 337.05 ± 26.7 μM) (Agusta et al., 2022). Methylations at position 4?′on the agathisflavone skeleton appear to be the least likely to suppress both cell activity, as the IC50 value of A2 was observed to be greater than 4. A comparison of the MCF-7 and HeLa cancer cells inhibitory activities of cupressuflavone derivatives (A1, 1, and 2) was shown to be a strong inhibitor of both cells MCF- 7 and HeLa cancer cells (IC50 between < 20 and 21–200 μM). On the contrary, agathisflavone derivatives (A2, and 4) were significantly less active (IC50 in ranging of 21–200 and 201–500 μM) (Damasuri et al., 2020).
Compound 3 belongs to the amentoflavone group, which has two methoxy groups at C4 and 4?′. It is very active in inhibiting the growth of both cell MCF-7 and HeLa cancer cells with IC50 values of 2.14 ± 0.6 and 11.03 ± 2.9 μM, respectively. The results of this study are slightly different from those of Li et al. (2019), who reported that 3 has an IC50 value of 8.38 ± 0.63 μM for HeLa cancer cells and 91.19 ± 0.5 μM for MCF-7 cells. The difference in the IC50 value is believed to occur due to differences in the condition (health) of the cancer cells used and the condition of the environment. Furthermore, Li et al. (2019) also reported that the amentoflavone derivative, 7,4′,4?′-tri-O-methyl amentoflavone with three methoxy groups shows proliferative activities on both cancer cells, respectively, the IC50 of 14.79 ± 0.64 μM for HeLa cancer cell and 57.62 ± 2.11 μM for MCF-7 cell. In general, the IC50 value is still very active for inhibiting HeLa cancer cells (IC50 < 20 μM), but it is less active in inhibiting MCF-7 cells (IC50 ranging from 21–200 μM). This suggests that the methoxy group at position C7 of the amentoflavone may have reduced its inhibitory effects on both cancer cells.
Compounds 1, 2, and 3 showed very active inhibition with the IC50 value of 11.54 ± 3.4, 3.40 ± 0.3, and 2.14 ± 0.6 μM, respectively. Compound 3 has the best activity against MCF-7 cells. In addition, compound 2 (IC50 of 1.42 ± 1.1 μM) having the best inhibitory activity against HeLa cancer cells, compared to 3 (IC50 11.03 ± 2.9 μM). All compounds have a higher IC50 value compared to epirubicin-HCl and napradox-50 (IC50 0.55 ± 0.2 and 0.35 ± 0.03 μM, respectively), indicating that all samples have lower activity than the positive controls.
The different substituents showed specific interactions between corresponding receptors/target molecules. As an anticancer agent, biflavonoids are known for their ability to control cell proliferation, apoptosis, invasion, metastasis, autophagy, transcription, and drug resistance in various types of cancers, including breast and cervical cancer cells. Biflavonoids may bind to the respective target and inhibit their activity to support cancer cells to grow. Moreover, the substituents attached also have an important role in controlling the energy of those interactions (Xiong et al., 2021).
Compound 2, which has a C8-C8? bond, would be the most potent inhibitor of the HeLa cancer cells (IC50 = 1.42 1.1 μM), in contrast to 3, having a C3′-C8? bond and is the strongest inhibitor of the MCF-7 cell (IC50 = 2.14 0.6 μM). Inhibitory studies using cupressuflavone and amentoflavone derivatives on proliferating cells are necessary to fully assess the anticancer potential of these drugs. Coulerie et al. (2013) reported that the robustaflavone, which has a C3′-C6? linkage, is a potent inhibitor of the DENV-NS5 RdRp (IC50 = 0.33 μM). However, the 7?-O-methylamentoflavone (sotetsuflavone), which has a C3′-C8? linkage, is the strongest inhibitor with IC50 = 0.16 μM. Based on SAR analysis, the skeleton of a biflavonoid provides a compelling template for the production of an anticancer compound. The number and position of -OCH3 groups, as well as the type of biflavonoid, all affect its anticancer effectiveness. Furthermore, a link was also discovered between patterns of oxygenation substitution and the control of the anticancer potential (Laishram et al., 2015; Tchimene et al., 2016; Wu et al., 2016). Biflavonoids are dimers of flavonoids through a C-O-C or C-C connection. The fundamental functional property of flavonoids is their capacity to act as antioxidants, which provide protection against cancer and heart disease. Their activity is strongly correlated with the position and number of hydroxyls that attached to their aromatic rings, the physicochemical properties of another substituent, and the capacity for intramolecular hydrogen bond formation (Çetinkaya et al., 2022; Scotti et al., 2013).
CONCLUSION
Five biflavonoids from the leaves of A. hunsteinii K. Schum. Indonesia have been successfully isolated. These compounds were identified as 7,4?′-di-O-methycupressuflavone (1), 7-O- methylcuppresuflavone (2), 4′, 4?′-di-O-methyl amentoflavone (3), 7,7?-di-O-methylagathisflavone (4) and 7,4′,7?,4?′-tetra-O-methyl-cupressuflavone (5). Compound 1 is the first isolated from the genus Araucaria. At the same time, 2-5 have been reported from several species of the genus Araucaria, but this is the first time it has been identified in A. husteinii. Regarding the SAR analysis, we also suggested that cupressuflavone and amentoflavone, are the most promising building block for developing anticancer drugs among biflavonoids. The number and location of methyl groups on the biflavonoid moiety modulate their inhibition growth of MCF-7 and HeLa cancer cells. A strong relationship was also discovered between patterns of oxygenation and the control of the anticancer potential. The most active compound and potent inhibitor against HeLa cancer cells are compounds 2 and 3, with an IC50 of 1.42 ± 1.1 and 11.03 ± 2.9 μM, respectively. Additionally, 1, 2, and 3 are also the most active compound and are the strongest inhibitor against MCF-7 cells with an IC50 of 11.54 ± 3.4, 3.40 ± 0.3, and 2.14 ± 0.6 μM, respectively.
LIST OF ABBREVIATIONS
CC, Column chromatography; DMSO, Dimethyl sulfoxide; ESI/MS, Electrospray ionization mass spectrometry; HMBC, Heteronuclear multiple bond coherence; HSQC, Heteronuclear singular quantum coherence; IR, Infrared; KBr, Potassium bromide; LC-MS/MS, Liquid chromatography-mass spectrometry tandem mass spectrometry; MCF-7, Michigan Cancer Foundation-7; MTT, [3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide; NCI, National Cancer Institute; NMR, Nuclear magnetic resonance; SAR, Structure-activity relationship; TLC, Thin layer chromatography; UV, Ultraviolet; VIS, Visible; WHO, World Health Organization.
ACKNOWLEDGMENTS
This project was funded by the Indonesian Ministry of Education, Culture, Research, and Technology through IPB University under Project No. 102/E5/PG.02.00.PL/2023, dated on June 19, 2023. In addition, the authors are grateful to the Primate Research Center (PSSP) IPB University and ITB Bandung for providing the platforms and facilities for this research.
AUTHOR CONTRIBUTIONS
The author contributions could be described as follows: Purwantiningsih Sugita conceived the design research, supported the analysis and research material, and revised the manuscript. Dhea Demitri Agusta, Kurniawanti, and Luthfan Irfana conducted the experiment, drafted, and revised the manuscript. Hanhan Dianhar and Dyah Utami Cahyaning Rahayu supervised the separation, structure elucidation, and revised the manuscript. Irma Herawati Suparto supervised the cytotoxicity sssay and revised the manuscript. The final manuscript has been reviewed and approved by all authors.
CONFLICTS OF INTEREST
The authors declare that there are no conflicts of interest.
ETHICAL APPROVALS
This study does not involve experiments on animals or human subjects.
DATA AVAILABILITY
All data generated and analyzed are included within this research article.
PUBLISHER’S NOTE
This journal remains neutral with regard to jurisdictional claims in published institutional affiliation
REFERENCES
Agusta DD, Dianhar H, Rahayu DUC, Suparto IH, Sugita P. Anticancer and antivirus activities of two biflavonoids from Indonesian Araucaria hunsteinii K Schum leaves. J Hunan Univ Nat Sci, 2022; 49(3): 168–77. doi: https://doi.org/10.55463/issn.1674-2974.49.3.18
Aslam MS, Choudhary BA, Uzair M, Subhan Ijaz A. Phytochemical and ethno- pharmacological review of the genus Araucaria—review. Trop J Pharm Res, 2013; 12:651–9. doi: https://doi.org/10.4314/tjpr.v12i4.31
Ayepola OR, Cerf ME, Brooks NL, Oguntibeju OO. Kolaviron, a biflavonoid complex of Garcinia kola seeds modulates apoptosis by suppressing oxidative stress and inflammation in diabetes-induced nephrotoxic rats. Phytomedicine, 2014; 21:1785–93. doi: https://doi.org/10.1016/j.phymed.2014.09.006
Çetinkaya S, Akça KT, Süntar I. Chapter 3—flavonoids and anticancer activity?: structure—activity relationship. Stud Nat Prod Chem, 2022; 74:81–115. doi: https://doi.org/10.1016/B978-0-323-91099-6.00017-7
Coulerie P, Nour M, Maciuk A, Eydoux C, Guillemot JC, Lebouvier N, Hnawia E, Leblanc K, Lewin G, Canard B, Figadère B. Structure-activity relationship study of biflavonoids on the dengue virus polymerase DENV-NS5 RdRp. Planta Med, 2013; 79:1313–8. doi: https://doi.org/10.1055/s-0033-1350672
Damasuri AR, Sholokhah EN, Mustofa. Cytotoxicity of ((E)-1-(4-aminophenyl)-3- phenylprop-2-en-1-one)) on HeLa cell line. Indones J Pharmacol Ther, 2020; 1:54–9. doi: https://doi.org/10.22146/ijpther.606
Deforest JC, Du L, Joyner PM. 4,4t′,7,7″-Tetra- O -methylcupressuflavone inhibits seed germination of Lactuca sativa. J Nat Prod, 2014; 77:1093–6. doi: https://doi.org/10.1021/np4010739
Frezza C, Venditti A, De Vita D, Toniolo C, Franceschin M, Ventrone A, Tomassini L, Foddai S, Guiso M, Nicoletti M, Bianco A, Serafini M. Phytochemistry, chemotaxonomy, and biological activities of the Araucariaceae family—a review. Plants, 2020; 9(7):1–73. doi: https://doi.org/10.3390/plants9070888
Haenel F, Garbow N. Cell counting and cnfluency analysis as quality controls in cell- based assays. Perkin Elmer Inc., Waltham, MA, 2014.
Hosseinzadeh E, Banaee N, Nedaie HA. Cancer and treatment modalities. Curr Cancer Ther Rev, 2017; 13:17–27. doi: http10.2174/1573394713666170531081818
[IARC] International Agency for Research on Cancer. IARC marks cervical cancer awereness Moth 2022 [WWW Document]. Available via https://www.iarc.who.int/featured-news/iarc-marks-cervical-cancer-awareness-month-2022/ (Accessed 03 October 2022).
Ilyas M, Seligmann O, Wagner H. Biflavones from the leaves of Araucaria rulei F. Muell. and a survey on biflavonoids of the Araucaria genus. J Zeitschrift für Naturforsch, 1977; 32(3–4): 206–9. doi: https://doi.org/10.1515/znc-1977-3-409
Khan NU, Ilyas M, Rahman W, Mashima T, Okigawa M, Kawano N. Biflavones from the leaves of Araucaria bidwilli Hooker and Agathis alba Foxworthy (Araucariaceae). Tetrahedron, 1972; 28:5689–95. doi: https://doi.org/10.1016/S0040-4020(01)88913-4
Laishram S, Sheikh Y, Moirangthem DS, Deb L, Pal BC, Talukdar NC, Borah JC. Anti-diabetic molecules from Cycas pectinata Griff. traditionally used by the Maiba-Maibi. Phytomedicine, 2015; 22:23–6. doi: https://doi.org/10.1016/j.phymed.2014.10.007
Li M, Li B, Xia ZM, Tian Y, Zhang D, Rui WJ, Dong JX, Xiao FJ. Anticancer effects of five biflavonoids from Ginkgo biloba L. male flowers in vitro. Molecules, 2019; 24:1–13. doi: https://doi.org/10.3390/molecules24081496
Menezes JCJMDS, Campos VR. Natural biflavonoids as potential therapeutic agents against microbial diseases. Sci Total Environ, 2021; 769:145168. doi: https://doi.org/10.1016/j.scitotenv.2021.145168
Molfetta FA, Honório KM, Alves CN, Da Silva ABF. A study on the anti-HIV activity of biflavonoid compounds by using quantum chemical and chemometric methods. J Mol Struct THEOCHEM, 2004; 674:191–7. doi: https://doi.org/10.1016/j.theochem.2003.12.007
Nakata R, Yoshinaga N, Teraishi M, Okumoto Y, Huffaker A, Schmelz EA, Mori N. A fragmentation study of isoflavones by IT-TOF-MS using biosynthesized isotopes. Biosci Biotechnol Biochem, 2018; 82:1309–15. doi: https://doi.org/10.1080/09168451.2018.1465810
Nawaz A, Jamal A, Arif A, Parveen Z. In vitro cytotoxic potential of Solanum nigrum against human cancer cell lines. Saudi J Biol Sci, 2021; 28:4786–92. doi: https://doi.org/10.1016/j.sjbs.2021.05.004
O’Brien K, Matlin AJ, Lowell AM, Moore MJ. The biflavonoid isoginkgetin is a general inhibitor of pre-mRNA splicing. J Biol Chem, 2008; 283:33147–54. doi: https://doi.org/10.1074/jbc.M805556200
Sasikala M, Sundaraganapathy R, Mohan S. MTT assay on anticancer properties of phytoconstituents from ipomoea aquatica forsskal using MCF–7 cell lines for breast cancer in women. Res J Pharm Technol, 2020; 13:1356–60. doi: https://doi.org/10.5958/0974-360X.2020.00250.4
Scotti L, Jaime Bezerra Mendonca Junior F, Rodrigo Magalhaes Moreira D, Sobral da Silva M, Pitta RI, Tullius Scotti M. SAR, QSAR and docking of anticancer flavonoids and variants: a review. Curr Top Med Chem, 2013; 12:2785–809. doi: https://doi.org/10.2174/1568026611212240007
Tchimene MK, Anaga AO, Ugwoke CEC, Onoja OJ, Ezugwu CO, Okunji C, Iwu MM. Anti-diabetic profile of extract, kolaviron, biflavonoids and garcinoic acid from Garcinia kola seeds. Int J Curr Microbiol Appl Sci, 2016; 5:317–22. doi: https://doi.org/10.20546/ijcmas.2016.502.036
Tsalikis J, Abdel-nour M, Farahvash A, Sorbara MT, Poon S, Philpott DJ. Isoginkgetin, a natural biflavonoid proteasome inhibitor, sensitizes cancer cells to apoptosis via disruption of lysosomal homeostasis and impaired protein clearance. Mol Cell Biol, 2019; 39(10):1–17. doi: https://doi.org/10.1128/MCB.00489-18
van Tonder VA, Joubert AM, Cromarty AD. Limitations of the 3-(4,5-dimethylthiazol-2- yl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) assay when compared to three commonly used cell enumeration assays. BMC Res Notes, 2015; 8:1–10. doi: https://doi.org/10.1186/s13104-015-1000-8
[WHO] World Health Organization. WHO launches new roadmap on breast cancer. 2023; [WWW Document]. Available via https://www.who.int/news/item/03-02-2023-who-launches-new-roadmap-on-breast-cancer (Accessed 17 March 2023).
Wu B, Song HP, Zhou X, Liu XG, Gao W, Dong X, Li HJ, Li P, Yang H. Screening of minor bioactive compounds from herbal medicines by in silico docking and the trace peak exposure methods. J Chromatogr A, 2016; 1436:91–9. doi: https://doi.org/10.1016/j.chroma.2016.01.062
Xie Y, Zhou X, Li J, Yao XC, Liu WL, Kang FH, Zou ZX, Xu KP, Xu PS, Tan GS. Identification of a new natural biflavonoids against breast cancer cells induced ferroptosis via the mitochondrial pathway. Bioorg Chem, 2021; 109:104744. doi: https://doi.org/10.1016/j.bioorg.2021.104744
Xiong X, Tang N, Lai X, Zhang J, Wen W, Li X, Li A, Wu Y, Liu Z. Insights into amentoflavone: a natural multifunctional biflavonoid. Front Pharmacol, 2021; 12:768708. doi: https://doi.org/10.3389/fphar.2021.768708
Yeap SK, Ali NM, Akhtar MN, Razak NA, Chong ZX, Ho WY, Boo L, Zareen S, Kurniawan TA, Avtar R, Ng SYL, Ong AHK, Alitheen NB. Induction of apoptosis and regulation ofMicroRNA expression by (2E,6E)-2,6-bis-(4-hydroxy-3-methoxybenzylidene)- cyclohexanone (BHMC) treatment on MCF-7 breast cancer cells. Molecules, 2021; 26:1–15. doi: https://doi.org/10.3390/molecules26051277
Yoon SO, Shin S, Lee HJ, Chun HK, Chung AS. Isoginkgetin inhibits tumor cell invasion by regulating phosphatidylinositol 3-kinase/Akt-dependent matrix metalloproteinase-9 expression. Mol Cancer Ther, 2006; 5:2666–75. doi: https://doi.org/10.1158/1535-7163.MCT-06-0321
Yu S, Yan H, Zhang L, Shan M, Chen P, Ding A, Li SFY. A review on the phytochemistry, pharmacology, and pharmacokinetics of amentoflavone, a naturally-occurring biflavonoid. Molecules, 2017; 22:1–23. doi: https://doi.org/10.3390/molecules22020299
Zhang YX, Li QY, Yan LL, Shi Y. Structural characterization and identification of biflavones in Selaginella tamariscina by liquid chromatography-diode-array detection/electrospray ionization tandem mass spectrometry. Rapid Commun Mass Spectrom, 2011; 25:2173–86. doi: https://doi.org/10.1002/rcm.5090