INTRODUCTION
The viral vector has been an effective vehicle and practical in clinics currently (Duan, 2018; Ong et al., 2019). Among nonviral vectors, polyethylenimine 25K (PEI25K) has been reported as an effective gene delivery carrier among cationic polymers (Jiang et al., 2017; Ryu et al., 2018). Modification of PEI has been reported to improve the transfection efficiency of the gene into the cells (Lisha Mali et al., 2021; Wu et al., 2020).
Plasmid DNA is a gene-based strategy. It has potential to be an alternative therapy for many diseases including cancer (Shimada, 2018), inherited retinal disease (Zhang et al., 2021), and cardiovascular disease (Gorabi et al., 2018).
Safe gene delivery is a vital issue that serves the vehicle to be effective and practical for clinical application and treatment (Clanchy et al., 2008; Ma et al., 2019). The objective of this current study was to investigate PEI25K-CL on the improved transfection efficiency of pEGFP and low cytotoxicity in HeLa cervical carcinoma cells.
MATERIALS AND METHODS
Materials
Polyethylenimine (branched, 25 KDa) was purchased from Sigma-Aldrich (St. Louis, MO). PEI 25K-CL (1:1 mole ratio) was provided by Dr. Boon-ek Yingyongnarongkul, Department of Chemistry and Center of Excellence for Innovation in Chemistry, Faculty of Science, Ramkhamhaeng University, Bangkok, Thailand. The XLarge DNA Ladder was purchased from GeneDireX (Las Vegas city, NV). pEGFP encoding enhanced green fluorescent protein was purchased from Clontech (Palo Alto, CA). HeLa human cervical carcinoma cells were obtained from the American Type Culture Collection (ATCC, Rockville, MD).
Plasmid DNA preparation
pEGFP was extracted and purified from DH5α Escherichia coli containing pEGFP using the Geneaid Plasmid Maxi Kit (Taipei City, Taiwan). The plasmid DNA was extracted according to the manufacturer’s instructions. The purified plasmid was dissolved in tris-ethylenediamine tetraacetic acid (EDTA) buffer, pH 8.0. DNA concentration was measured at 260 nm using a cuvette photometer (BioPhotometer, Eppendorf AG, Hamburg, Germany).
Preparation of PEI25K-CL/pEGFP complexes
PEI25K-CL (0.1 mg/ml) in sterile water and pEGFP (1 mg/ml) in tris-EDTA buffer, pH 8.0, were diluted in ultrapure water or in Minimum Essential Medium (MEM). The PEI25K-CL and pEGFP complexes based on weight ratios of 0.125:1 to 2.5:1 were formed by adding a pEGFP solution to a PEI25K-CL solution, pipetted up-down for 15 times, and allowing spontaneous complex assembling for 15–20 minutes at room temperature (Weecharangsan et al., 2021).
Measurement of buffer capacity of PEI 25K-CL
PEI 25K-CL (1:1 molar ratio) was dissolved in ultrapure water at a concentration of 1 mg/ml and then diluted with 10 ml of a 0.9% sodium chloride solution to a concentration of 0.1 mg/ml. The PEI 25K-CL solution was then titrated with a 0.1 N hydrochloric acid solution of 0.5–8.0 μmol. Simultaneously, the pH of the PEI 25K-CL solution was monitored using a pH meter (Bench 700, Oakton Instrument, IL). The buffer capacity of PEI 25K-CL was calculated by dividing the amount of hydrochloric acid by pH change at a high resistance of pH change (Sinko, 2006).
Gel electrophoresis assay
To analyze whether pEGFP was compacted by PEI25K-CL, a gel retardation assay was performed by electrophoresis (Weecharangsan et al., 2021). The PEI25K-CL/pEGFP complexes at polymer/DNA ratios of 0:1, 0.125:1, 0.25:1, 0.5:1, 0.75:1, and 1:1 were prepared and loaded with loading dye Invitrogen (Grand Island, NY) into the wells of 1% agarose gel prepared in tris-borate-EDTA buffer containing 0.74% SYBR Green. The sample-loaded gel was electrophoresed at the voltage of 100 V for 15–20 minutes. The DNA bands were visualized on ImageQuant LAS 4000 Mini (GE Healthcare Bioscience AB, Upsala, Sweden).
Particle size, polydispersity index, and zeta potential
The particle size, polydispersity index, and zeta potential of the PEI25K-CL/pEGFP complexes in ultrapure water were determined using the Zetasizer Nano Series (Malvern Instruments Ltd., Worcestershire, UK) at the temperature of 25°C.
In vitro transfection and pEGFP expression
HeLa cells were plated in a 24-well plate at a density of 2 × 104 cells/well with MEM + 10% fetal bovine serum (FBS) for 24 hours. 1 μg/well of pEGFP was gently mixed with 0.5–2.5 μg/well of PEI25K-CL and incubated for 15 minutes at room temperature. The PEI25K-CL/pEGFP complexes were subjected to HeLa cell transfection for 4 hours, and the transfected cells were rinsed with phosphate buffered saline (PBS) pH 7.4 and incubated with a complete growth medium at 37°C for 24 hours. The LipofectamineTM2000/pEGFP complexes at liposome/DNA ratio of 0.92/1 (v/w) were used as control. The cells were observed under a fluorescence microscope (Eclipse TS100, Nikon, Tokyo, Japan). The number of GFP-expressed cells was counted. The transfection efficiency was defined as the ratio of GFP expressed cell number to the plate area (Paecharoenchai et al., 2012).
Cytotoxicity of PEI25K-CL
HeLa cells were seeded at a density of 8 × 103 cells/well in a 96-well plate in MEM containing 10% FBS and grown for 24 hours. PEI25K-CL and PEI25K at the concentration of 0 to 20 μg/ml were added to each well and incubated for 4 hours. Cells treated with a growth medium were used as control. After 4 hours, the cells were then rinsed with PBS pH 7.4 and incubated with a growth medium for 24 hours. The 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide assay was used to detect the cytotoxicity of PEI25K-CL (Weecharangsan et al., 2014).
Statistical analysis
The data were collected as mean ± SD. The significant difference in the data was analyzed using Student’s t-test and analysis of variance as p < 0.05.
RESULTS
Buffer capacity of PEI25K-CL
Buffer capacity of PEI25K-CL and PEI25K was calculated at the high region resistance of pH change 7.12 ± 0.05 to 7.76 ± 0.04 and 7.01 ± 0.07 to 7.77 ± 0.12, respectively, at the amount of HCl of 1.5 × 10−3 μmol (Fig. 1). The buffer capacity of PEI25K-CL and PEI25K was 0.0023 ± 0.0002 and 0.002 ± 0.0002, respectively. The buffer capacity of PEI25K-CL was not different from that of PEI25K. However, the amount of HCl needed for neutralization of PEI25K was higher than that of PEI25K-CL for the pH range of 7.0–7.7. This could be due to the amount of free NH2 group of PEI25K being higher than that of PEI25K-CL.
Electrophoresis of PEI25K-CL/pEGFP complexes
Gel electrophoresis was used to examine the PEI25K-CL/pEGFP complexes and PEI25K/pEGFP complexes (Fig. 2). The electrophoretic mobility of pEGFP of the PEI25K-CL/pEGFP and PEI25K/pEGFP complexes was completely retained at the polymer/DNA ratios of above 0.5:1. At the polymer/DNA ratio of 0.5:1, the PEI25K-CL/pEGFP complexes exhibited higher DNA band intensity than that of the PEI25K/pEGFP complexes. This suggested that PEI25K-CL had a lower ability to form a complex with pEGFP than that of PEI25K.
Colloidal properties of PEI25K-CL/pEGFP complexes
Figure 3 shows the particle size, polydispersity index, and zeta potential of the PEI25K-CL/pEGFP and PEI25K/pEGFP complexes performed in ultrapure water. The particle size, polydispersity index, and zeta potential of the PEI25K-CL/pEGFP and PEI25K/pEGFP complexes were 220 ± 7 to 402 ± 34 and 271 ± 12 to 462 ± 8 nm, 0.39 ± 0.05 to 0.51 ± 0.16 and 0.47 ± 0.07 to 0.6 ± 0.07, and 32.9 ± 1.4 to 43.7 ± 1.7 and 37.2 ± 4.6 to 43.5 ± 1.8 mV, respectively.
The particle size, polydispersity index, and zeta potential of the PEI25K-CL/pEGFP and PEI25K/pEGFP complexes were dependent on the polymer/DNA ratio. At the polymer/DNA ratios of 0.5:1 and 2.5:1, the particle size of the PEI25K-CL/pEGFP complexes was significantly larger than that of the PEI25K/pEGFP complexes, while at the polymer/DNA ratios of 0.5:1 and 2.5:1 the particle size of the PEI25K-CL/pEGFP complexes was significantly smaller than that of the PEI25K/pEGFP complexes. At the polymer/DNA ratio of 0.5:1, the polydispersity index and zeta potential of the PEI25K-CL/pEGFP complexes were significantly lower than those of the PEI25K/pEGFP complexes while not significantly different at other polymer/DNA ratios.
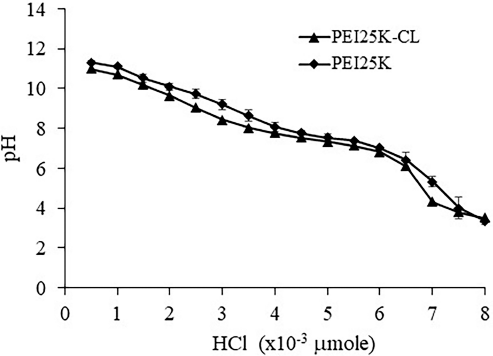 | Figure 1. Buffer capacity of PEI25K-CL and PEI25K analyzed by acid-base titration. [Click here to view] |
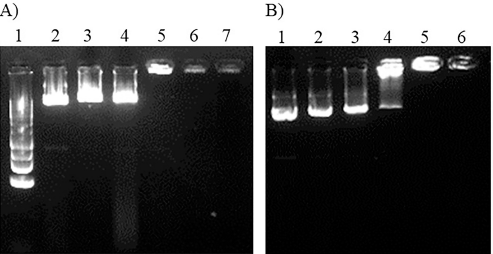 | Figure 2. gel electrophoresis of PEI25K-CL/pEGFP complexes and PEI25K/pEGFP complexes on 1% agarose gel A) DNA ladder: lane 1, PEI25K/pEGFP complexes at polymer/DNA ratios of 0:1, 0.125:1, 0.25:1, 0.5:1, 0.75:1 and 1:1; lane 2-7, respectively (15 min electrophoresis); B) PEI25K-CL/pEGFP complexes at polymer/DNA ratios of 0:1, 0.125:1, 0.25:1, 0.5:1, 0.75:1 and 1:1; lane 1-6, respectively (20 min electrophoresis). [Click here to view] |
pEGFP expression of PEI25K-CL/pEGFP and transfection efficiency
pEGFP expression of the PEI25K-CL/pEGFP and PEI25K/pEGFP complexes in HeLa cells analyzed by fluorescent microscopy is shown in Figure 4. PEI25K-CL/pEGFP exhibited lower pEGFP expression than that of PEI25K-CL/pEGFP. Transfection efficiency of pEGFP into HeLa cells of the PEI25K-CL/pEGFP complexes was 8 ± 4 to 679 ± 295 cells/cm2 depending on the polymer/DNA (Fig. 5A). The transfection efficiency of pEGFP into HeLa cells of the PEI25K-CL/pEGFP complexes was significantly higher than in cells treated with pEGFP (Fig. 5A) while significantly lower than in cells treated with the PEI25K/pEGFP complexes and LipofectamineTM2000/pEGFP complexes (Fig. 5B).
Cytotoxicity of PEI25K-CL
The cytotoxicity of PEI25K-CL and PEI25K in HeLa cells is shown in Figure 6. The cell viability of PEI25K-CL and PEI25K was 23.6% ± 0.8% to 99.5% ± 1.5% and 20.0% ± 0.9% to 97.2% ± 3.4%, respectively. The cell viability of PEI25K-CL and PEI25K was not different at the polymer concentration of 0.1 μg/ml, while that of PEI25K-CL was significantly lower than that of PEI25K at the polymer concentration of all above 0.1 μg/ml.
DISCUSSION
Our study demonstrated that PEI25K-CL had the ability to deliver plasmid DNA and yielded an efficient transfection efficiency and low cytotoxicity. The low cytotoxicity and efficient transfection efficiency of PEI25K-CL could be due to the amine groups of PEI25K partially substituted by cholesterol (Weecharangsan et al., 2021).
 | Figure 3. Particle size (A), polydispersity index (B) and zeta potential (C) of PEI25K-CL/pEGFP and PEI25K/pEGFP complexes analyzed at polymer/DNA ratios of 0.5:1, 1:1, 2:1 and 2.5:1. *, significantly different. [Click here to view] |
 | Figure 4. pEGFP expression of PEI25K-CL/pEGFP (A) and PEI25K/pEGFP (B) complexes in HeLa cells at polymer/DNA ratio of 1:1 analyzed by fluorescent imaging. [Click here to view] |
 | Figure 5. GFP expressed HeLa cells A) PEI25K- CL/pEGFP complexes at polymer/DNA ratios of 0.5:1, 1:1, 1:1.5, 2:1 and 2.5:1. *, significantly different from cells treated with pEGFP; B) PEI25K-CL/pEGFP complexes and PEI25K/pEGFP complexes at polymer/DNA ratio of 1:1, and LipofectamineTM2000/pEGFP complexes at liposome/DNA ratio of 0.92/1 (v/w). *, significantly different from cells treated with PEI25K-CL/pEGFP complexes. [Click here to view] |
PEI25K-CL had a buffer capacity similar to that of PEI25K and a lower neutralizing ability of HCl at the pH range of 7.12 ± 0.05 to 7.76 ± 0.04. Wang et al. (2002) showed that PEI1.8K-CL and PEI10K-CL had a high buffer capacity in the pH range of 5–7. Weecharangsan et al. (2021) showed that the buffer capacity of PEI25K-CA was not different from that of PEI25K.
PEI25K-CL had the ability to condense plasmid DNA, pEGFP. Jiang et al. (2010) exhibited that polyethylenimine–cholesterol cationic lipopolymer could condense plasmid DNA. Wang et al. (2002) showed that PEI1.8K-CL and PEI10K-CL could condense DNA to the nanosized and positively charged complexes.
 | Figure 6. Cytotoxicity of PEI25K-CL and PEI25K at polymer concentration of 0.1 to 20 μg/ml in HeLa cells. *, significantly different. [Click here to view] |
The colloidal property often dictates the cellular entry mechanism of carrier/DNA complexes. The PEI25K-CL/pEGFP complexes yielded nanosized and positively charged particles. Li et al. (2018) demonstrated that polyethylenimine–cholesterol/miR was positively charged and nanosized.
PEI25K-CL improved pEGFP expression and transfection efficiency and low cytotoxicity in HeLa oral carcinoma cells. Jiang et al. (2010) exhibited that polyethylenimine–cholesterol lipopolymer/DNA had a good transfection efficiency in A549 and MCF-7 cells. Wang et al. (2002) showed that the PEI1.8K-CL/pCMS-EGFP and PEI10K-CL/pCMS-EGFP complexes had high GFP expression in Jurkat cells. Remant et al. (2020) demonstrated that cholesterol-grafted low-molecular-weight PEI enabled siRNA delivery into K562 myeloid leukemia cells depending on the degree of substitution of PEI backbone. Sun et al. (2014) reported that PEI-cholesterol liposomes improved cellular uptake and endosomal escape of cisplatin, as did Wu et al. (2020) in hydrophobization of PEI with cholesterol of siRNA.
PEI25K-CL had low cytotoxicity in HeLa oral carcinoma cells. Gusachenko et al. (2009) showed that lipophilic conjugated-PEI25K had low cytotoxicity at particular modification of amino groups of the polymer. Jiang et al. (2010) exhibited that the polyethylenimine–cholesterol cationic lipopolymer/DNA complexes were low cytotoxic in A549 and MCF-7 cells. Wu et al. (2020) exhibited that hydrophobization of PEI with cholesterol was low cytotoxic compared with PEI. Wang et al. (2002) showed that PEI1.8K-CL and PEI10K-CL had slight cytotoxicity in Jurkat cells. Sun et al. (2014) exhibited that PEI-cholesterol liposomes did not show noticeable systemic toxicity in H22 hepatoma-bearing mice. Li et al. (2018) demonstrated that polyethylenimine–cholesterol was nontoxic in CAL-27 cells at the concentration of 0–20 μg/ml. Lin et al. (2017) demonstrated that PEI25K induced autophagy in mouse fibroblast cells. Anwer et al. (2013) showed that PEG-cholesterol lipopolymer/plasmid IL-12 did not exhibit worsened side effects in the chemotherapy treatment of a phase I trial.
CONCLUSION
This study concluded that PEI25K-CL had potential to be an efficient and safe gene delivery carrier.
AUTHORS’ CONTRIBUTIONS
WW and BY contributed to the concept and design. WW contributed to the data acquisition and interpretation and the writing and final approval of the manuscript. PO contributed to the research supervision.
CONFLICTS OF INTEREST
The authors have no conflicts of interest to declare.
FUNDING
This study was supported by the Faculty of Pharmacy, Srinakharinwirot University (034/2556).
ETHICAL APPROVALS
This study does not involve experiments on animals or human subjects.
DATA AVAILABILITY
All data generated and analyzed are included within this research article.
PUBLISHER’S NOTE
This journal remains neutral with regard to jurisdictional claims in published institutional affiliation.
REFERENCES
Anwer K, Kelly FJ, Chu C, Fewell JG, Lewis D, Alvarez RD. Phase I trail of a formulated IL-12 plasmid in combination with carboplatin and docetaxel chemotherapy in the treatment of platinum-sensitive recurrent ovarian cancer. Gynecol Oncol, 2013; 131:169–73. CrossRef
Clanchy FIL, Williams RO. Plasmid DNA as a safe gene delivery vehicle for treatment of chronic inflammatory disease. Expert Opin Biol Ther, 2008; 8:1507–19. CrossRef
Duan D. Systemic AAV micro-dystrophin gene therapy for Duchenne muscular dystrophy. Mol Ther, 2018; 26:2337–56. CrossRef
Gorabi AM, Hajighasemi S, Tafti HA, Soleimani M, Panahi Y, Ganjali S, Sahebkar A. Gene therapy in cardiovascular disease: a review of recent updates. J Cell Biochem, 2018; 119:9645–54. CrossRef
Gusachenko SO, Kravchuk Y, Konevets D, Silnikov V, Vlassov VV, Zenkova MA. Transfection efficiency of 25-kDa PEI-cholesterol conjugates with different levels of modification. J Biomater Sci Polym Ed, 2009; 20:1091–110. CrossRef
Jiang B, He H, Yao Li, Zhang T, Huo J, Sun W, Yin L. Harmonizing the intracellular kinetics toward effective gene delivery using cancer cells-targeted and light-degradable polyplexes. Biomacromolecules, 2017; 18:877–85. CrossRef
Jiang YN, Mo HY, Chen JH. Gene transfer system mediated by polyethylenimine-cholesterol lipopolymer with lipid microbubbles. Yao Xue Xue Bao, 2010; 45:659–66.
Lisha Mali A, Priya SS, Rekha MR. Hydrophobic and hydrophilic modifications of polyethylenimine towards gene delivery applications. J Appl Polym Sci, 2021; 138:51323. CrossRef
Li L, Li X, Huang X, Jiang W, Liu L, Hou C, Yang Y, Zhang L, Zhang X, Ye L, Yuan J, Li G, Sun H, Mao L. Synergistic anticancer effects of nanocarrier loaded with berberine and miR-122. Biosci Rep, 2018; 38:BSR20180311. CrossRef
Lin CW, Jan MS, Kuo JS. The vector-related influences of autophagic microRNA delivery by Lipofectamine 2000 and polyethylenimine 25K on mouse embryonic fibroblast cells. Eur J Pharm Sci, 2017; 101:11–21. CrossRef
Ma Y, Wise AK, Shepherd RK, Richardson RT. New molecular therapies for the treatment of hearing loss. Pharmacol Ther, 2019; 200:190–209. CrossRef
Ong T, Pennesi ME, Birch DG, Lam BL, Tsang SH. Adeno-associated viral gene therapy for inherited retinal disease. Pharm Res, 2019; 36:34. CrossRef
Paecharoenchai O, Niyomtham N, Apirakaramwong A, Ngawhirunput T, Rojanarata T, Yingyongnarongkul B, Opanasopit P. Structure relationship of cationic lipids on gene transfection mediated by cationic liposomes. AAPS PharmSciTech, 2012; 13:1302–8. CrossRef
Remant KC, Thapa B, Valencia-Serna J, Domun S, Dimitroff C, Jiang X, Uluda?. Cholesterol-grafted low cationic lipopolymers: potential siRNA carriers for selective chronic myeloid leukemia therapy. J Biomed Mater Res A, 2020; 108:565–80. CrossRef
Ryu N, Kim MA, Park D, Lee B, Kim YR, Kim KH, Baek JI, Kim WJ, Lee KY, K UK. Effective PEI-mediated delivery of CRISPR-Cas9 complex for targeted gene therapy. Nanomedicine, 2018; 14:2095–12. CrossRef
Shimada H. P53 molecular approach to diagnosis and treatment of esophageal squamous cell carcinoma. Ann Gastroenterol Surg, 2018; 2:266–73. CrossRef
Sinko PJ. Buffer capacity. In: Martin’s physical pharmacy and pharmaceutical sciences. Lippincott Williams & Wilkins, New York, NY, pp 213−7, 2006.
Sun X, Chen J, Gu X, Liang W, Wang J. Efficacy and toxicity of cisplatin liposomes modified with polyethylenimine. Pharmazie, 2014; 69:281–6.
Wang D, Narang AS, Kotb M, Gaber AO, Miller DD, Kim SW, Mahato RI. Novel branched poly(ethylenimine)-cholesterol water-soluble lipopolymers for gene delivery. Biomacromolecules, 2002; 3:1197–207. CrossRef
Weecharangsan W, Paecharoenchai O, Niyomtham N, Opanasopit P, Yingyongnarongkul B, Lee RJ. Cholic acid-modified polyethelenimine for gene delivery into human carcinoma cells. Adv Mat Res, 2014; 1060:3–6. CrossRef
Weecharangsan W, Niyomtham N, Yingyongnarongkul B, Opanasopit P, Lee RJ. Growth inhibition of human carcinoma cells by cholic acid-conjugated polyethylenimine 25K/p53-EGFP complexes on human carcinoma cells. J Appl Pharm Sci, 2021; 11:14–21.
Wu P, Luo X, Wu H, Zhang Q, Wang K, Sun M, Oupicky D. Combined hydrophobization of polyethylenimine with cholesterol and perfluorobutyrate improves siRNA delivery. Bioconjug Chem, 2020; 31:698–707. CrossRef
Zhang D, McLenachan S, Chen SC, Zaw K, Alziyadat Y, Zhang X, Lamey TM, Thompson JA, McLaren TL, Mellough C, De Roach JN, Chen FK. Generation of two induced pluripotent stem cell lines from a patient with recessive inherited retinal disease caused by compound heterozygous mutations in SNRNP200. Stem Cell Res, 2021; 51:102154. CrossRef