INTRODUCTION
Bacteria are microorganisms consisting of Gram-positive and Gram-negative. Gram-positive bacteria are composed of a peptidoglycan wall containing a specific surface polysaccharide and having polar properties, whereas Gram-negative bacteria have three layers of cell, namely, outer membrane, cell wall, and inner plasma membrane so that antibacterial compounds will be more difficult to inhibit Gram-negative bacterial activity (Glauert and Thornley, 1969). Some bacteria can cause serious infections such as endocarditis (Holland et al., 2016), meningitis (Wenger et al., 1990), pneumonia (Dasaraju and Liu, 1996), and sepsis (Minasyan, 2019).
Chlorine, the most common disinfectant, is moderately oxidative and reacts with various components of bacterial cells (Kim et al., 2008) so that the presence of Cl halogen atoms can play a role in inactivating various microorganisms such as Escherichia coli, Enterococcus faecalis, Psedomonas aeruginosa, Bacillus subtilis, and S. aureus but remain safe for humans because chlorine has lower toxin levels compared to other halogen compounds such as fluorine and bromine (McDonnell and Russell, 1999). In addition to the presence of Cl atoms, heterocyclic rings play a role in medicinal chemistry as therapeutic agents including acting as antitumor (Chen et al., 2014; El-Sawy et al., 2013), antifungal (Cao et al., 2014; Russell and Soiket, 2015), antiviral (Salem et al., 2013; Stamatiou et al., 2003), anti-inflammatory (El-Sawy et al., 2014), and broad spectrum of antibacterial drugs (El-Sawy et al., 2013; Mustafa, 2018).
Hexagamavunon-6 (HGV-6) is a curcumin analog which was synthesized by the Faculty of Pharmacy, Gadjah Mada University, and has been known to have active antibacterial activity on Gram-positive bacteria with 1,000 μg/ml inhibitory concentrations (Sardjiman, 2000). The activity of HGV-6 predicted because the structure has four chloro and two hydroxy groups in which plays a role in denaturing proteins in the bacterial cell wall.
The objective of this research was to synthesis analogs curcumin carried out by reacting 2-chlorobenzaldehyde, 4-chlorobenzaldehyde, and 2,4-dichlorobenzaldehyde with N-methyl-4-piperidone which is a heterocyclic compound with a nitrogen atom that still has one pair of free electrons. All of the synthesized compounds tested for antibacterial activity against Gram-positive and Gram-negative bacteria using the Agar diffusion method. Each compound has a chloro atom in an aromatic group and N atom in a heterocyclic group of N-methyl-4-piperidone.
To assist in identifying molecular targets from the active side of the compound structure, the chemical docking method is used (Meng et al., 2011). Structural models of proteins in bacteria will be inhibited by compounds obtained through Protein Data Banks (PDB).
MATERIALS AND METHODS
Materials
All starting materials, reagents, and solvents were of reagent grade for analysis. Synthesis was controlled using UV lamp 254 and 366 nm (Desaga Heidelberg) with Thin Layer Chromatography (TLC) silica gel F254 (Merck). Melting point was recorded by Buchi melting point B540, Elucidation 0structure of the compound was recorded on Fourier-transform infrared spectroscopy (FTIR) Perkin Elmer and beam splitter KBr/Gemid-infrared, Direct Inlet-Mass Spectrometry (DI-MS) (QP2010S SHIMADZU), 1H-NMR (JOEL), and 13C-NMR (JOEL).
General procedure for synthesis
N-methyl-4-piperidone (1 mmol) was stirred with sodium hydroxide (0,1 gram) as a catalyst. Then aromatic aldehydes (2 mmol) were added as the starting material, and ethanol (2 ml) as the solvent was added dropwise until it was dissolved. The stirring reaction is carried out until the starting material is not left over, controlled by using TLC. The reaction is stopped when the TLC plate has formed a single product spot and then the mixture was washed with aquadest and dried at 45°C for 24 hours (Gregory et al., 2013).
 | Scheme 1. Basic frameworks. [Click here to view] |
In vitro antimicrobial activity assay
The antibacterial activities were tested using the agar diffusion method. Bacterial culture (Streptococcus mutans, E. coli ATCC 25922, B. subtilis ATCC 6633, P. aeruginosa ATCC 27853, Klebsiella Pneumoniae, and E. faecalis ATCC 29212) was prepared in Brain Heart Infusion broth and adjusted with Optical Density at a wavelenght of 600 nm to 108 cfu/ml (CLSI, 2012). Mueller Hintor Agar was poured on a different sterile petri dish and allowed to solidify. Then, the surface of the agar medium was inoculated with a sterile swab with the bacterial strain. All test samples were prepared in Dimethyl sulfoxide (DMSO) (2 mg/ml) and 10 μl of each concentration added in a paper disk. DMSO was used as negative control, and amoxicilin (31.25 μg/ml) was used as positive control in all experiments. Finally, all petri dishes were incubated for 24 hours at 37°C. The results of antibacterial activity were evaluated as diameter inhibiton. It was determined by the inhibition zone diameter on test plates against the compound and compared with control antibiotic inhibition zone (Balouiri et al., 2016).
Molecular modeling studies
Preparation of ligand structure
All of the compounds (1a-c), HGV-6 and Amoxicillin, the reference molecules of the docking simulations, were built in the two-dimensional model in the form of Molecular Operating Environment (MOE) database. The determination of conformation was then performed on the database. Furthermore, its minimal energy was calculated (Wijianto et al., 2019).
Docking simulations
MOE was used to evaluate the interaction between the compound and target protein of curcumin analogs so that it can inhibit the bacterial activity. The target protein was obtained from the PDB (ID code: 5OJ0). 5OJ0 is a protein for the bacterium Streptococcus pneumoniae. The bacterium is a Gram-positive bacterium whose bacterial cell wall structure can be used as a reference to see interactions in the cell wall structure of other Gram-positive bacteria. Protein contains a reference molecule that is Cefixime as the native ligand and group of Penicillin-Binding Protein 2X (PBP2X). All of the docking conformations were analyzed, and the best value with the precise pose was selected for further interaction study.
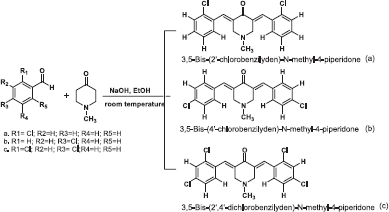 | Scheme 2. Reagent and conditions of synthesis (Gregory et al., 2013). [Click here to view] |
RESULTS AND DISCUSSION
Synthesis
Synthesis of 3,5-Bis-(2'-chlorobenzilyden)-N-methyl-4-piperidone (a)
Yellow crystal, yield = 72%, m.p. = 145°C
IR (KBr, cm−1) 3425,58 (N-CH3 streatching); 3062,96 (=C-H streatchcing aromatic); 2939,52 (C-H streatching aliphatic); 1674,21 (C=O streatching aromatic); 1627,92 (C=C streatching aliphatic); 1597,06 (C=C streatching aromatic); 756,10 (subtitued ortho).
1HNMR δ 2,347 (3H, s,N-CH3); 3,588 (4H, s, CH2); 7,207 (2H, m, C-H aromatic); 7,267 (4H, m, C-H aromatic); 7,415 (2H, m, C-H aromatic); 7,979 (2H, s, C-H alkene)
13CNMR δ 45,756 (R2-N-CH3); 56,881 (R2-N-CH2-C-R); 126,625 (C-aromatic); 130,148 (C-aromatic); 130,215 (C-aromatic); 130,541(C-aromatic); 133,814 (C-aromatic); 134,217 (Cα unsaturated); 134,448 (Cβ unsaturated); 135,369 (C-Cl); 186,358 (C=O)
DI-MS (EI, 70 eV) 100% area with [M+1] 357
Synthesis of 3,5-Bis-(4'-chlorobenzilyden)-N-methyl-4-piperidone (b)
Pale yellow crystal, yield = 44%, m.p. = 170°C
IR (KBr, cm−1) 3448,72 (N-CH3 streatching); 3032,10 (=C-H streatching aromatic); 2939,52 (C-H streatching aliphatic); 1674,21 (C=O streatching carbonil); 1627,92 (C=C streatching aliphatic); 1581,63 (C=C streatching aromatic); 825,53 (subtitued para).
1HNMR δ 2,444 (3H, s,N-CH3); 3,701 (4H, d, CH2); 7,292 (4H, m, C-H aromatic); 7,367 (4H, m, C-H aromatic); 7,723 (2H, s, C-H alkene).
13CNMR δ 45,121 (R2-N-CH3); 57,178 (R2-N-CH2-C-R); 129,092 (C-aromatic); 131,799 (C-aromatic); 133,584 (C-aromatic); 133,786 (Cα unsaturated); 135,340 (Cβ unsaturated); 135,398 (C-Cl); 186,713 (C=O).
DI-MS (EI, 70 eV) 100% area with [M+1] 357
3,5-Bis-(2',4'-dichlorobenzilyden)-N-methyl-4-piperidone (c)
Yellow crystal, yield = 60%, m.p = 151°C
IR (KBr, cm−1) 3425,58 (N-CH3 streatching); 3070,66 (=C-H streatching aromatic); 2939,52 (C-H streatching aliphatic); 1689,64 (C=O streatching carbonil); 1612,49 (C=C streatching aliphatic); 1581,63 (C=C streatching aromatic).
1HNMR δ 2,356 (3H, s,N-CH3); 3,548 (4H, s, CH2); 7,136 (2H, d, C-H aromatic); 7,153 (2H, d, C-H aromatic); 7,456 (4H, d, C-H aromatic); 7,889 (2H, s, C-H alkene).
13CNMR δ 45,813 (R2-N-CH3); 56,814 (R2-N-CH2-C-R); 127,095 (C-aromatic); 130,119 (C-aromatic); 133,584 (C-aromatic); 131,194 (C-aromatic); 132,259 (C-aromatic); 133,142 (Cα unsaturated); 134,755 (Cβ unsaturated); 135,571 (C-Cl); 136,166 (C-Cl); 185,945 (C=O).
DI-MS (EI, 70 eV) 100% area with [M+1] 427
Antibacterial activity assay
Synthesized compounds were screened for in vitro activity by determining the inhibitory diameter against three Gram-positive and three Gram-negative bacteria and compared with the standard antibiotic, amoxicillin, by the Agar diffusion method. The inhibitory diameter of microbes is seen from the presence of clear areas formed around the disk paper which can then be measured using a ruler in millimeters (mm). DMSO solvent is used to dissolve the synthesized compound with a concentration of 100% and also used as a negative control. The positive control used is the antibiotic amoxicillin for the antibacterial test which is a class of β-lactam antibiotics (Akhavan and Vijhani, 2019). The amoxicillin concentration used is 31.25 μg/ml. After incubation at 37°C for 24 hours, the zone of inhibition of growth around each disk was measured and zone diameters were interpreted. The experiment was performed in triplets, and the results of the antibacterial activity test can be seen in Table 1, which is the result of observations of the inhibition diameter of bacterias using synthesized compounds by analog curcumin 1a-c.
Most of the synthesized compounds have inhibition on the Gram-positive and Gram-negative bacteria at a concentration of 1,000 μg/ml except K. pneumonia in compound 1b. Compound 1c was found to have significant inhibition in both Gram-positive and Gram-negative bacteria up to a concentration of 125 μg/ml. Compounds 1a and 1b display moderate antimicrobial activity in Gram-positive bacteria and weak antimicrobial activity in Gram-negative bacteria. The result of antibacterial activity shows that the compound 1c has four chloro substitution on a benzene ring carrying electron-withdrawing groups which have three free electron pairs so that it can form more hydrogen bonds with amino acids in bacterial cells compared to compounds 1a-b which only have two chlorine atoms. The inhibition of S. mutans, B. subtilis, and E. faecalis bacteria, which are a class of Gram-positive bacteria, has a greater average inhibition diameter compared to E. coli, P. aeruginosa, and K. pneumoniae bacteria which are a Gram-negative bacterium. Gram-positive bacteria are composed of peptidoglycan wall containing a specific surface polysaccharide and have polar properties. Whereas Gram-negative bacterias have three layers of cell wrapping, namely, outer membrane, cell wall, and inner plasma membrane so that antibacterial compounds will be more difficult to inhibit Gram-negative bacteria activity (Silhavy et al., 2010).
To further investigate the molecular inhibitory activity of compounds in bacteria, an in silico study was conducted using docking. Docking results showed S values in Table 2, whereas the S value is the Gibbs free energy to measure the strength of the bond between the ligand with the site of action on the target protein. The smaller the S value, the stronger the bond formed with the ligand. Compound 1c has the S value that closest to the positive control, amoxicillin.
Figure 1 shows the docking interaction of amoxicillin with the two amino acids Asn on bacterial cell wall through hydrogen bonds that are formed in the lone pair of electrons from the nitrogen atom in β-lactam structure and in the NH2 group. Figure 2 shows the docking interaction of compound 1c with one Asn amino acid in bacteria cell wall through nitrogen atoms in heterocyclic groups, N-methyl-4-piperidone. So that compound 1c has the best activity compared to other synthesis compounds but has not been able to be better than Amoxicillin.
 | Table 1. In vitro antibacterial activity of the compound tested by Agar diffusion method as the diameter (mm) of the inhibition zone. [Click here to view] |
 | Table 2. S value of docking results from each test compound [Click here to view] |
 | Figure 1. Docking interaction 5Oj0 with Amoxicillin. [Click here to view] |
 | Figure 2. Docking interaction 5Oj0 with 1c compound. [Click here to view] |
CONCLUSION
All of the compounds can be synthesized using condensation reaction and produce moderate to excellent yields. The synthesized compound was evaluated for their in vitro antibacterial activity and indicated that all of the compounds have inhibitory activity in Gram-positive and Gram-negative bacteria at concentrations of 1,000 μg/ml except K. pneumonia in compound 1b. The 3,5-Bis-(2',4'-dichlorobenzilyden)-N-methyl-4-piperidone (1c) compound has the most significant inhibition activity which is Gram-positive bacteria up to a concentration of 62.5 μg/ml and Gram-negative bacteria to a concentration of 125 μg/m. Compound 1c has the potential to be used as a drug with antibacterial activities through interaction and PBP2X inhibition confirm by docking studies.
FINANCIAL SUPPORT
The authors would like to thank to Kementerian Riset dan Teknologi Pendidikan Tinggi for the research grant of Penelitian Berbasis Kompetensi (PBK) 2018 and Penelitian Dasar (PD) 2019.
CONFLICT OF INTEREST
The authors declare that they have no conflict of interest.
REFERENCES
Akhavan BJ, Vijhani P. Amoxicillin. Treasure Island, FL: StatPearls Publishing, 2019.
Balouiri M, Sadiki M, Ibnsouda SK. Methods for in vitro evaluating antimicrobial activity: a review. J Pharm Anal, 2016; 6:71–9. CrossRef
Cao X, Sun Z, Cao Y, Wang R, Cai T, Chu W. Design, synthesis, and structure-activity relationship studies of novel fused heterocycles-linked triazoles with good activity and water solubility. J Med Chem, 2014; 57:3687–706. CrossRef
Chen Y, Yu K, Tan NY, Qiu RH, Liu W, Luo NL. Synthesis, characterization and anti-proliferative activity of heterocyclic hypervalent organoantimony compounds. Eur J Med Chem, 2014; 79:391–8. CrossRef
Clinical & Laboratory Standarts Institute (CLSI). M02-A11: performance standards for antimicrobial disk susceptibility tests; Approved standard (11th Edition). Wayne, Pensylvania: Clinical and Laboratory Standards Institute, 2012.
Dasaraju PV, Liu C. Infections of the respiratory system: medical microbiology. Galveston, TX: University of Texas Medical Branch, 1996.
El-Sawy ER, Ebaid MS, Abo-Salem HM, Al-Sehemi AG, Mandour AH. Synthesis, anti-inflammatory, analgesic and anticonvulsant activities of some new 4,6-dimethoxy-5-(heterocycles)benzofuran starting from naturally occurring visnagin. Arab J Chem, 2014; 7:914–23. CrossRef
El-Sawy ER, Mandour AH, El-Hallouty SM, Shaker KH, Abo-Salem HM. Synthesis, antimicrobial and anticancer activities of some new N-methylsulphonyl and N-benzenesulphonyl-3-indolyl heterocycles: 1st Cancer Update. Arab J Chem, 2013; 6:67–78. CrossRef
Glauert AM, Thornley MJ. The topography of the bacterial cell wall. Annu Rev Microbiol, 1969; 23:159–98. CrossRef
Gregory M, Dandavati A, Lee, Megan, Tzou S, Savagian M, Brien KA. Synthesis, cytotoxicity, and structure–activity insight of NH- and N-methyl-3,5-bis-(arylidenyl)-4-piperidones. Med Chem Res, 2013; 22:5588–97. CrossRef
Holland TL, Baddour LM, Bayer AS, Hoen B, Miro JM, Fowler VG. Infective endocarditis. Nat Rev Dis Primers, 2016; 2:16059. CrossRef
Kim J, Pitts B, Stewart PS, Camper A, Yoon J. Comparison of the antimicrobial effects of chlorine, silver ion, and tobramycin on biofilm. Antimicrob Agents Chemother, 2008; 52:1446–53. CrossRef
McDonnell G, Russell AD. Antiseptics and disinfectants: activity, action, and resistance. Clin Microbiol Rev, 1999; 12:147–79. CrossRef
Meng XY, Zhang HX, Mezei M, Cui M. Molecular docking: a powerful approach for structure-based drug discovery. Curr Comput-Aid Drug, 2011; 7:146–57. CrossRef
Minasyan H. Sepsis: mechanisms of bacterial injury to the patient. SJTREM, 2019; 27:19. CrossRef
Mustafa YF. Synthesis, characterization and antibacterial activity of novel heterocycle, coumacine, and two of its derivatives. SJP, 2018; 26:870–5. CrossRef
Russell M, Soiket MIH. Some new azole type heterocyclic compounds as antifungal agents. Org Communications, 2015; 7:114–22.
Salem MS, Sakr SI, El-Senousy WM, Madkour HMF. Synthesis, antibacterial, and antiviral evaluation of new heterocycles containing the pyridine moiety. Arc Pharm, 2013; 346:766–73. CrossRef
Sardjiman S. Synthesis of some new series of curcumin analouge, antioxidative, antiinflammatory, antibacterial activities, qualitative structure-activity relationship (Dissertation). Yogyakarta, Indonesia: Universitas Gadjah Mada, 2000.
Silhavy TJ, Kahne D, Walker S. The bacterial cell envelope. CSH Perspect Biol, 2010; 2:a000414. CrossRef
Stamatiou G, Foscolos GB, Fytas G, Kolocouris A, Kolocouris N, Pannecouque C. Heterocyclic rimantadine analogues with antiviral activity. Bioorg Med Chem, 2003; 11:5485–92. CrossRef
Wenger JD, Hightower AW, Fecklam RR, Gaventa S, Broome CV. Bacterial meningitis in the United States. J Infect Dis, 1990; 162:1316–23. CrossRef
Wijianto B, Ritmaleni, Purnomo H, Nurrochmad A. In silico and in vitro assay of HGV analogue as antibacterial. Int J Pharm Pharm Sci, 2019; 78–85. CrossRef