INTRODUCTION
Hepatitis C virus (HCV) is the major causative agent of Hepatitis C disease. Its infection leads to chronic liver diseases, such as hepatitis, cirrhosis, and hepatocellular carcinoma or liver failure. It is the most common endemic liver diseases worldwide affecting approximately 71 million people (WHO, 2017). In viral hepatitis, hepatocytes cells are replaced by tough and non-functional fibrous tissue (Clin, 2009). The HCV is a micro RNA virus belonging to a member of the Flaviviridae family. Its genetic material is the ribonucleic acid (RNA) which encodes the structural proteins (Core proteins E1 and E2) and the nonstructural proteins include NS3 protease and NS5B RNA dependent RNA polymerase (Thayaillany et al., 2015). NS5B protein has been considered as a target to design and develop noble anti-HCV agents. The nonstructural protein 5B (NS5B), an RNA dependent RNA polymerase, is responsible for the replication of HCV genome (Varun et al., 2014). It has been clinically tested that NS5B plays a central role in the HCV replication. Most of the current drugs target this enzyme for the treatment of hepatitis C diseases. Hence, NS5B enzyme is considered as a target to design and develop a potential drug against HCV. However, the present antihepatitis drug treatment is expensive, less effective, and has numerous side effects. Thus, specific antiviral agents need to be developed against HCV life cycle.
Medicinal plants have been considered as an alternative source of medicines for the treatment of various human diseases, including liver diseases (Kumar and Pandey, 2013). Experimental studies showed that medicinal plants possess secondary metabolites known as phytochemicals. Many phytochemicals are being tested against hepatitis virus to find the promising antiviral agents. They have been reported to have many medicinal activities, such as anti-oxidant, anti-inflammatory, anti-thrombogenic, anti-mutagenic, anti-allergic, anti-bacterial, anti-cancer, antiviral, neuroprotective, and hepatoprotective (Panche et al., 2016; Shakya, 2016).
Therefore, the aim of the present study is to investigate medicinal plants derived phytochemicals as NS5B inhibitors against hepatitis C NS5B polymerase in silico approach.
MATERIALS AND METHODS
Ligand preparation
Ligands selected for the protein (NS5B) interactions were based on review of the literature on potential hepatoprotective phytochemicals derived from medicinal plants. Medicinal plants derived phytochemicals along with the reference ligand, Dasabuvir are summarized in Table 1. The chemical structure of ligands was accessed from the PubChem database and drawn in Advanced Chemistry Development’s Chemsketch which were converted into 3D structure (Programme Database file-PDB) by using the Open Babel software and employed for docking analysis. The Dasabuvir, a known antihepatitis drug, was used as a reference ligand to compare the docking results of phytochemicals (Trivella et al., 2015).
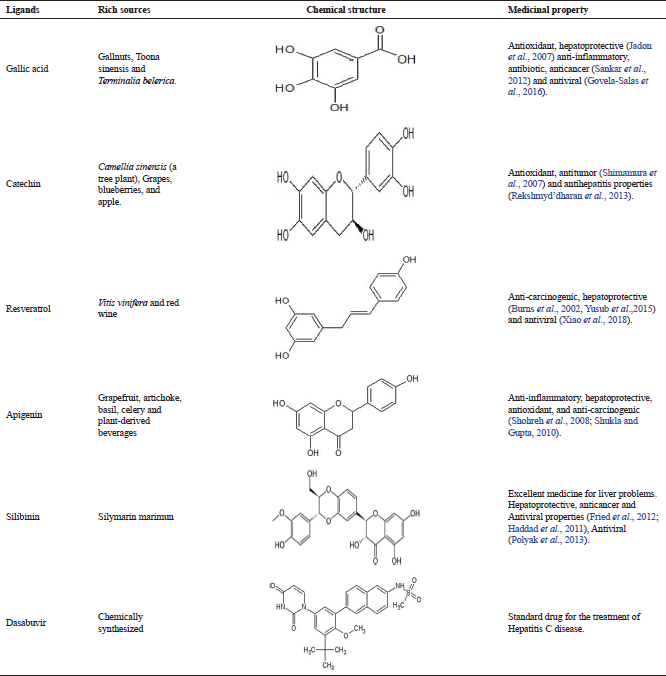 | Table 1. Chemical structures of ligands constructed using ChemSketch Software. [Click here to view] |
Druglikeness calculation
Molinspiration (http://www.molinspiration.com/) is an online tool used to determine the drug-likeness properties of ligands. Lipinski's Rule of Five is a rule of thumb to evaluate druglikeness properties. It states that most “drug-like” molecules should have logP ≤ 5, molecular weight ≤ 500, number of hydrogen bond acceptors ≤ 10, and number of hydrogen bond donors ≤ 5.
Preparation of protein structure
The crystallographic structure of the NS5B polymerase bound 4, 5-dihydrofurano indole was retrieved from the protein data bank (PDB ID: 3UPI) and the resolution was 2.00 Å (Fig. 1) (Velazquez et al., 2011). The 3UPI protein contains two types of polypeptide chains: A and B. The bound inhibitor 4, 5-dihydrofurano indole (HETATM and CONNECT) was removed from the 3UPI. The protein structure was then optimized using Swiss-PDB viewer software. The optimized structure has minimum energy confirmation which was used for the docking study.
Active site prediction
The NS5B active site was predicted by the online server of Meta Pocket 2.0 (Huang, 2009). The potential amino acid residues at NS5B active site was given in Table 2.
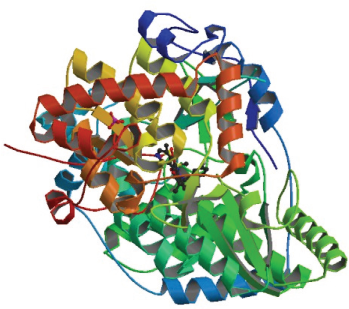 | Figure 1. HCV NS5B Polymerase (PDB ID: 3UPI). [Click here to view] |
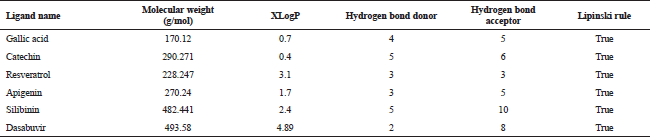 | Table 2. Druglikeness properties of ligands evaluated using the molinspiration tool. [Click here to view] |
Docking studies
The docking of all the phytochemicals toward the active site of the 3UPI (NS5B) targets was conducted using the AutoDock 4.2 tool (Morris et al., 2009). During the AutoDock, all the water molecules were removed, the hydrogen atoms were added to the 3UPI on which atom force field charges and atom types were assigned. The ligand (phytochemicals) and the receptor protein NS5B (3UPI) were first converted from PDB format into PDBQT format by Autodock tool. To run autodock, the grid box parameter values, Kollman charges and atomic solvation parameters were added. Grid maps dimensions were set as 60 × 60 × 60 points with spacing of 0.469A° to yield the receptor model that included atoms within 0.5A° of the grid centre. After Grid generation and efficient search of torsional freedom, the ligands were docked with the receptor 3UPI (NS5B). The Lamarckian genetic was used as a scoring algorithm. The number of runs for each docking experiment was set to 10. The remaining parameters were set as default. The protein-ligands docked complex was further evaluated on lowest binding energy (kcal/mol) values using the Root-mean-square-deviation data. The result of docked ligand-receptor binding interactions was visualized using the Discovery Studio 2019 client.
RESULTS AND DISCUSSION
The molecular docking is a computational tool which used to predict the binding affinity between ligands and target receptor proteins (Kumar and Doss, 2016). In the present study, druglikeness properties of all the phytochemicals were determined first and then docked with the HCV NS5B polymerase in silico approach. Traditional medicinal plants have also been used for treating diverse viral diseases for centuries. A number of natural phytochemicals experimentally proved that they have an antiviral activity against various kinds of viruses (Mathew et al., 2014; Sangeetha and Rajarajan, 2015). Balavignesh et al. (2013) reported that NS5B polymerase is an RNA dependent RNA polymerase enzyme which plays an important role in replication of HCV.
The druglikeness scores of the selected ligands were analyzed using the Molinspiration tool (http://www.molinspiration.com) as per Lipinski’s rule which determines whether particular molecule is similar to the known drugs (Lipinski et al., 2012, Mishra et al., 2018). It is an important parameter to determine four simple physicochemical parameter ranges (MW ≤ 500, log P ≤ 5, Hbond donors ≤ 5, H-bond acceptors ≤ 10) (Arumugam et al., 2012). The molecular weight of all the selected ligands was in the acceptable range (MW ≤ 500). Compounds with low molecular weight can be easily absorbed, distributed, and transported as compared to high molecular weight compounds. The XLogP value is used to assess the lipophilic efficiency that determines the potency of drug. The Molecular weight (MW), XLogP, H-bond donors, and H-bond acceptors value of all the phytochemicals were calculated and found to be within acceptable range according to Lipinski’s rule (Table 2).
 | Table 3. Potential ligand binding sites in NS5B active site (3UPI) obtained from the MetaPocket 2.0. [Click here to view] |
Table 3 revealed that the ligand binding site in the NS5B protein was predicted using the MetaPocket 2.0 on which all the five ligands confirmed the most conserved interacting amino acids of NS5B compared to the reference ligand, Dasabuvir. According to the Huang and Schroeder (2008), protein binding site prediction helps to improve protein docking. The present study revealed that there are 14 interactive amino acids residues involved between the NS5B receptor protein and the ligands. These interacting residues include GLU B:143, SER B:367, ARG B:386, ARG B:394, SER B:368, ASN B:411, SER B:407, TYR B:448, CYS B:366, SER B:470, LEU B: 384 ASN B:316, PRO B:197, and LEU B:384 constituting the potential binding active site for the HCV-NS5B. The docking score, binding amino acid residues, H-bond distance (Å), and number of hydrogen bonds are given in Table 4. All the ligands were interacted with the type B polypeptide chain of NS5B which was used in the docking study.
Figure 2 depicted the docking complex between the Gallic acid and the NS5B. Gallic acid formed five H-bonds with the residues of NS5B, include GLU143, SER367, and ARG386. The docking score was −5.48 kcal/mol. Figure 3 showed the binding relationship between the catechin and the NS5B with the binding energy −7.31 kcal/mol. It makes four hydrogen bonds with the amino acids SER368, SER407, TYR448, and ASN411 in the binding pocket of NS5B active site. The docked resveratrol-NS5B protein complex is formed two hydrogen bonding with the amino residues TYR448, CYS 366 of the NS5B as shown in the Figure 4. It has binding energy −8.14 kJ/mol. Figure 5 exhibited the hydrogen bonding interactions between the Apigenin and the NS5B which docks one amino acid SER 470 at NS5B active site and its docking score was −8.75 kcal/mol. Figure 6 demonstrates the docking complex between the silibinin and the NS5B with the binding energy −10.57 kcal/mol. This docking showed three H-bond interactions with the amino acid residues: LEU384 and ASN316 at NS5B binding site. The reference ligand, dasabuvir showed high docking score −11.43 kcal/mol and bound to the residue PRO197 and LEU384 of NS5B binding site via three H-bonds (Fig. 7). The binding energies of all the ligands phytochemical compounds were compared to the reference ligand (Dasabuvir). Therefore, these results demonstrated that there are good binding interactions between phytochemicals and NS5B polymerase. Based on minimum docking score, the most active ligands with the NS5B are catechin (−7.31 kcal/mol), resveratrol (−8.14 kcal/mol), apigenin (−8.75 kcal/mol), and silibinin (−10.57 kcal/mol) compared to the gallic acid (−5.48 kcal/mol). Therefore, these results suggested that silibinin has best binding score compared to other docked phytochemicals score compared to other docked ligands. With this good docking score, silibinin was proved to be most active compounds which may possess antihepatitis activity against HCV similar to the docked dasabuvir because both have hydrogen binding interactions with the LEU384 residues of NS5B active site. Silibinin has been tested against HCV and its antiviral effect was found to be effective in inhibiting the enzymatic activity of NS5B RNA-dependent RNA polymerase (Usman and Sobia, 2014). Our findings are also supported the antiviral effect of silibinin.
Kortemme et al. (2003) demonstrated that minimum the binding energy, greater is the binding affinity in silico study. In silico study also suggested that greater the hydrogen bond interactions between the enzyme (receptor protein) and ligand determines the strength of binding (Biswal et al., 2019). Elfiky (2019) reported that docking calculations are used to predict the binding affinity of the compounds to HCV NS5B active site (Kumar and Doss, 2017). The active functional group of phytochemicals are completely surrounded by the binding pocket of the NS5B because they fit properly into its active site via hydrogen bonding interactions (Jin et al., 2014). From the results of Docking analysis, it was found that all phytochemicals have antiviral inhibitory activity against HCV NS5B polymerase enzyme.
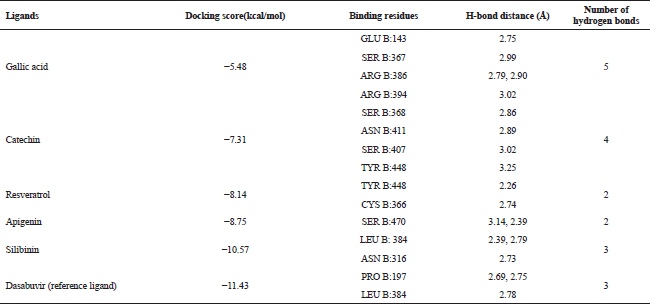 | Table 4. Molecular docking analysis of ligands against NS5B polymerase (3UPI). [Click here to view] |
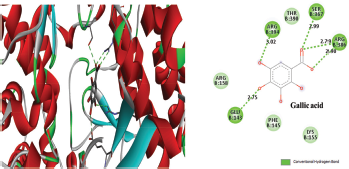 | Figure 2. Three-dimensional and two-dimensional interactions of Gallic acid against the active site of NS5B polymerase. [Click here to view] |
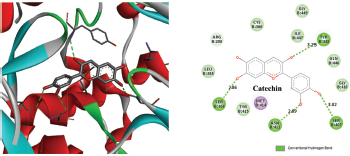 | Figure 3. Three-dimensional and two-dimensional interactions of Catechin against the active site of NS5B polymerase. [Click here to view] |
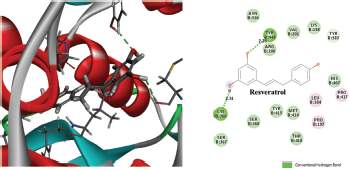 | Figure 4. Three-dimensional and two-dimensional interactions of Resveratrol against the active site of NS5B polymerase. [Click here to view] |
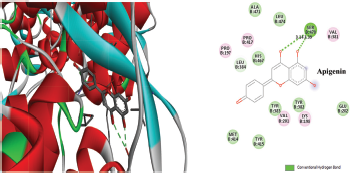 | Figure 5. Three-dimensional and two-dimensional interactions of Apigenin against the active site of NS5B polymerase. [Click here to view] |
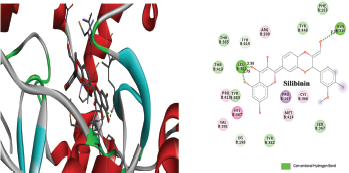 | Figure 6. Three-dimensional and two-dimensional interactions of Silibinin against the active site of NS5B polymerase. [Click here to view] |
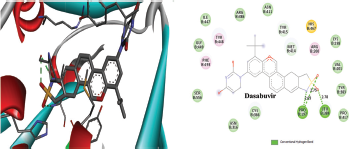 | Figure 7. Three-dimensional and two-dimensional interactions of Dasabuvir against the active site of NS5B polymerase. [Click here to view] |
CONCLUSION
This study suggested that all the phytochemical compounds could be potent HCV NS5B inhibitors which might be due to their good binding efficacy with the HCV NS5B which may further investigate in vitro and in vivo model.
ACKNOWLEDGMENTS
The author would like to thank the School of Sciences, Indira Gandhi National Open University, Maidan Garhi New Delhi-110068 India to carry out this research work.
CONFLICT OF INTEREST
The author confirms no conflict of interest for this manuscript.
FINANCIAL SUPPORT
None.
REFERENCES
Arumugam M, Muthuswamy U, Kuppusamy AK, Thirumalaisamy S, Varadharajan S, Puliyath J. Computational drug discovery of potential aldose reductase inhibitors using in silico studies. Electronic J Biol, 2012; 8:67–72.
Balavignesh V, Srinivasan E, Ramesh Babu NG. and Saravanan N. Molecular docking study on NS5B polymerase of hepatitis c virus by screening of volatile compounds from Acacia concinna and ADMET prediction. Int J of Pharm Life Sci, 2013; 4:2548–58.
Biswal AR, Venkataraghavan R., Vivek P, Ivo RS. Molecular docking of various bioactive compounds from essential oil of Trachyaspermum ammi against the fungal enzyme Candidapepsin-1. J Appl Pharma Sci, 2019; 9:021–32. CrossRef
Burns J, Yokota T, Ashihra H, Lean MEJ, Crozier A. Plant foods and herbal sources of resveratrol. J Agric Food Chem, 2002; 50:3337–40. CrossRef
Clin CJ. The global surden of Hepatitis C working group. Global Burden of Disease (GBD) for hepatitis C. Pharmacology, 2009; 44:20–9. CrossRef
Elfiky AA. Novel guanosine derivatives as anti-HCV NS5b polymerase: a QSAR and molecular docking study. Med Chem, 2019; 15:130–7. CrossRef
Fried MW, Navarro VJ, Afdhal N, Belle SH, Wahed AS, Hawke RL, Doo E, Meyers CM, Reddy KR. Effect of silymarin (milk thistle) on liver disease in patients with chronic hepatitis C who failed interferon therapy: a randomized, placebo-controlled trial. JAMA, 2012; 308:274–82. CrossRef
Govela-Salas M, Rivas-Estilla AM, Rodríguez-Herrera R, Lozano-Sepúlveda SA, Aguilar-Gonzalez CN, Zugasti-Cruz A, Salas-Villalobos TB, Morlett-Chávez JA. Gallic acid decreases hepatitis C virus expression through its antioxidant capacity. Exp Ther Med, 2016; 11:619–24. CrossRef
Haddad Y, Vallerand D, Brault A, Haddad PS. Antioxidant and hepatoprotective effects of silibinin in a rat model of nonalcoholic steatohepatitis. Evid Based Complement Altern Med, 2011; Article ID:647903:1–10. CrossRef
Huang B. MetaPocket: a meta approach to improve protein ligand binding site prediction. Omics, 2009; 13:325–30. CrossRef
Huang B, Schroeder M. Using protein binding site prediction to improve protein docking. Gene, 2008; 422:14–21. CrossRef
Jadon A, Bhadauria M, Shukla S. Protective effect of Terminalia belerica Roxb. and gallic acid against carbon tetrachloride induced damage in albino rats. J Ethnopharmacol, 2007; 109:214–8. CrossRef
Jin G, Lee S, Choi M, Son S, Kim GW, Oh JW, Lee C, Lee K. Chemical genetics-based discovery of indole derivatives as HCV NS5B polymerase inhibitors. Eur J Med Chem, 2014; 75:413–25. CrossRef
Kortemme T, Morozov AV, Baker D. An orientation-dependent hydrogen bonding potential improves prediction of specificity and structure for proteins and protein-protein complexes. J Mol Biol, 2003; 326:1239–59. CrossRef
Kumar DT, Doss CG. Investigating the inhibitory effect of Wortmannin in the hotspot mutation at Codon 1047 of PIK3CA kinase domain: a molecular docking and molecular dynamics approach. Adv Protein Chem Struct Biol, 2016; 102:267–97. CrossRef
Kumar DT, Doss CG. Role of E542 and E545 missense mutations of PIK3CA in breast cancer: a comparative computational approach. J Biomol Struct Dyn, 2017; 35:2745–57. CrossRef
Kumar S, Pandey A. Chemistry and biological activities of flavonoids: an overview. Sci World J, 2013; Article ID 162750:1–16. CrossRef
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Delivery Rev, 2012; 64:4–17. CrossRef
Mathew S, Faheem M, Archunan G, Ilyas M, Nargis B, Syed J, Ishtiaq Q, Qahtani. In silico studies of medicinal compounds against hepatitis C capsid protein from North India. Bioinform Biol Insights, 2014; 8:159–68. CrossRef
Mishra SS, Gaur R, Gajaram S, Kumar N, Sharma CS, Singh HP. Computational analysis of pharmacokinetic, bioactivity and toxicity parameters of some selected oral-hypoglycaemic agents. Chem Res J, 2018; 3:135–40.
Molinspiration Cheminformatics. Molinspiration. Available via http://www.molinspiration.com/cgi-bin/properties, 2016.
Morris, GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ. Autodock4 and AutoDockTools4: automated docking with selective receptor flexiblity. J. Comput Chem, 2009; 16:2785–91. CrossRef
Panche AN, Diwan A D, Chandra SR. Flavonoids: an overview. J Nutr Sci, 2016; 5:e47. CrossRef
Polyak SJ, Ferenci P, Pawlotsky JM. Hepatoprotective and antiviral functions of silymarin components in hepatitis C virus infection. Hepatology, 2013; 57:1262–71. CrossRef
Rekshmyd’dharan S, Roy A. Epicatechin nature’s extraordinary therapeutic agent: a review. Int J PharmTech Res, 2013; 5:1816–22.
Sangeetha Vani G, Rajarajan S. A study on in-silico analysis of phytochemicals targeting the proteins of hepatitis B and C virus. Int J Curr Microbiol App Sci, 2015; 4:683–91.
Sankar K, Tamizhmani T, Rama Krishna S, Krishna KVVS, Syam Vardhan M. Simultaneous estimation of gallic acid and rutin in marketed polyherbal formulations by HPTLC. Int J Pharmtech Res, 2012; 4:616–22.
Shakya AK. Medicinal plants: future source of new drugs. Int J Herbal Med, 2016; 4:59–64.
Shimamura T, Zhao WH, Hu ZQ. Mechanism of action and potential for use of tea catechin as an antiinfective agent. Antiinfect Agents Med Chem, 2007; 6:57–62. CrossRef
Shukla S, Gupta S. Apigenin: a promising molecule for cancer prevention. Pharm Res, 2010; 27:962–78. CrossRef
Thayaillany R, Radha P, Prabhu M. Molecular docking studies of indole derivatives containing cyanide group as hepatitis C Ns5b polymerase inhibitor. Pharma Innovation J, 2015; 4:43–8.
Trivella JP, Gutierrez J, Martin P. Dasabuvir: a new direct antiviral agent for the treatment of hepatitis C. Expert Opin Pharmacother, 2015; 16:617–24. CrossRef
Usman AA, Sobia I. Medicinal plants against hepatitis C virus. World J Gastroenterol, 2014; 20:2941–7. CrossRef
Varun G, Lokesh M, Sandeep M, Sajad Shahbazi, Deepak Reddy G. Novel indole derivatives as hepatitis C virus NS5B polymerase inhibitors: pharmacophore modeling and 3D QSAR studies. Bangladesh J Pharmacol, 2014; 9:290–7. CrossRef
Velazquez F, Venkataraman, S. Lesburg CA. Duca JS, Rosenblum SB, Kozlowski JA, Njoroge FG. Synthesis of novel 4,5-dihydrofurano indoles and their evaluation as HCV NS5B polymerase inhibitors. Org Lett, 2011; 14:556–559. CrossRef
WHO. Global hepatitis report, vol. 4, WHO, Geneva, Switzerland, pp 1–83, 2017.
Xiao Q, Zhu W, Feng W, Lee SS, Leung AW, Shen J, Gao L, Xu C. A review of resveratrol as a potent chemoprotective and synergistic agent in cancer chemotherapy. Front Pharmacol, 2018; 9:1534. CrossRef
Yusub A, Hassim H, Hamzah H, Noordin MM. Antiviral activity of resveratrol against human and animal viruses. Adv Virol, 2015; Article ID:184241:1–7. CrossRef